Abstract
Patients with Coronavirus disease 2019 exhibit low expression of interferon-stimulated genes, contributing to a limited antiviral response. Uncovering the underlying mechanism of innate immune suppression and rescuing the innate antiviral response remain urgent issues in the current pandemic. Here we identified that the dimerization domain of the SARS-CoV-2 nucleocapsid protein (SARS2-NP) is required for SARS2-NP to undergo liquid–liquid phase separation with RNA, which inhibits Lys63-linked poly-ubiquitination and aggregation of MAVS and thereby suppresses the innate antiviral immune response. Mice infected with an RNA virus carrying SARS2-NP exhibited reduced innate immunity, an increased viral load and high morbidity. Notably, we identified SARS2-NP acetylation at Lys375 by host acetyltransferase and reported frequently occurring acetylation-mimicking mutations of Lys375, all of which impaired SARS2-NP liquid–liquid phase separation with RNA. Importantly, a peptide targeting the dimerization domain was screened out to disrupt the SARS2-NP liquid–liquid phase separation and demonstrated to inhibit SARS-CoV-2 replication and rescue innate antiviral immunity both in vitro and in vivo.
Similar content being viewed by others
Main
Coronavirus disease 2019 (COVID-19), caused by the novel severe acute respiratory syndrome (SARS) coronavirus 2 (SARS-CoV-2), emerged and spread rapidly worldwide, posing a major threat to global public health and safety1. The SARS-CoV-2 genomic RNA is harboured within a viral particle consisting of four structural proteins—the crown-like spike (S) glycoprotein, the membrane (M) protein, the ion channel envelope (E) protein and the nucleocapsid protein (NP; SARS2-NP). The S protein binds to the human ACE2 receptor and is responsible for viral entry into host cells. The M protein facilitates viral assembly in the endoplasmic reticulum and the NP assembles with viral RNA to form a helical ribonucleoprotein complex called the nucleocapsid2,3.
Although the precise interaction between the innate immune system and SARS-CoV-2 is yet to be fully elucidated, it has been suggested that the innate immune response plays a crucial role in determining the severity of COVID-19 (refs. 4,5,6). During coronavirus infection, the viral components and replication intermediates serve as pathogen-associated molecular patterns recognized by host germline-encoded pattern recognition receptors, including retinoic acid-inducible gene I (RIG-I)-like receptor and members of the Toll-like receptor family7,8,9,10. The activation of innate immune receptors promotes the production of type I interferons (IFN-I; including IFN-β and IFN-α), which mediates the antiviral response through the upregulation of IFN-stimulated genes (ISGs)11,12,13,14,15.
The NP of coronaviruses associates with the large positive-strand RNA viral genome and the carboxy (C)-terminal endodomain of the M protein, which is critical for coronavirus replication16. SARS2-NP possesses self-assembly properties, with its dimerization domain (DD) being responsible for homodimer formation as well as C-terminal-mediated tetramer formation17. However, whether SARS2-NP possesses a pathogenic mechanism other than the packaging of viral RNA and whether the self-assembly properties of SARS2-NP could be targeted for COVID-19 prevention and therapy remain unknown. Here we reveal the mechanisms of NP-mediated innate immune evasion, NP modification and mutation, which together permit the optimization of therapeutic strategies and antiviral drug against COVID-19.
Results
The SARS2-NP DD is required for LLPS and viral RNA association
SARS2-NP contains three predicted intrinsically disordered regions (IDRs), an amino (N)-terminal RNA-binding domain and a C-terminal DD17,18,19 (Extended Data Fig. 1a). Given that proteins with IDRs tend to undergo liquid–liquid phase separation (LLPS)20,21, we tested whether NP has such a property. Purified green fluorescent protein (GFP)-conjugated NP (Extended Data Fig. 1b) formed microsized droplets in solutions (Fig. 1a) and more numerous and larger droplets were observed at low salt concentrations, weak acidic conditions (pH 4.5–7.5) and high temperatures (Fig. 1a–c). The droplets dissolved when the NaCl concentration was increased, suggesting that electrostatic interactions are needed (Fig. 1a). SARS2-NP droplet formation was inhibited by 5% 1,6-hexanediol (Hex)—a compound that disrupts weak hydrophobic interactions22 (Extended Data Fig. 1c)—and completely disrupted by heat or proteinase K (Extended Data Fig. 1d,e). The droplet size of NPs labelled with AF488 increased with time (Fig. 1d), indicating that the puncta were not artificially formed by the GFP tag. A fluorescence recovery after photobleaching (FRAP) assay revealed rapid fluorescence recovery after bleaching (Fig. 1e), indicating efficient diffusion within the SARS2-NP droplets.
a–c, Droplet formation (left) and phase separation (PS) diagram (right) of GFP-labelled SARS2-NP (5 μM) at room temperature with the indicated concentrations of NaCl at pH 5.5 (a), 150 mM NaCl at the indicated pH conditions (b) and at the indicated temperatures with 150 mM NaCl at pH 5.5 (c). d, Fusion and equivalent diameter (EqDiameter) of AF488-labelled SARS2-NP (5 μM) droplets with 150 mM NaCl in pH 5.5 at room temperature at the indicated time points. e, Representative micrographs of GFP–SARS2-NP droplets before and after photobleaching (left). FRAP quantification of GFP–SARS2-NP droplets over a period of 35 s (right; n = 3 droplets). f, Fusion of GFP–SARS2-NP condensates in a HeLa cell. g, Representative micrographs (left) and FRAP quantification (right) of GFP–SARS2-NP condensates in vivo before and after photobleaching (red arrow, bleach site) over a period of 35 s (n = 3 condensates). h, Immunofluorescence microscopy of HeLa (left) and U2OS (right) cells transfected with GFP–SARS2-NP and Cy5–RNA. i, In vivo fusion of Cy3–RNA and GFP–SARS2-NP condensates. j, Representative micrographs of droplet formation (left), and quantification of purified GFP–SARS2-NP WT and GFP–SARS2-NP that was depleted of its IDRs (d_IDR1, d_IDR2 and d_IDR3), RNA-binding domain (d_RBD) and DD domain (d_DD; right; 5 μM; n = 6 independent experiments). k, GFP–SARS2-NP WT, treated with or without 5% Hex for 30 s, and d_DD condensates in HeLa cells (left). Percentages of cells harbouring NP condensates (right; n = 3 independent experiments). l, Immunoblot (IB) of the total cell lysate (TCL) and immunoprecipitates (IP) derived from HEK293T cells transfected with SARS2-NP WT or its truncated plasmids. m, In situ PLA for Myc- and Flag-tagged SARS2-NP in HeLa cells (left). The PLA-detected proximity (PROX) complexes were quantified (right; n = 3 independent experiments). n, SPR assay of the binding affinity between RNA and SARS2-NP WT (left) and d_DD (right). Different SARS2-NP concentrations (1–5 nM) were used to calculate the association rate (Ka), the dissociation constant (Kd), and equilibrium dissociation constant (KD) values. r.u., resonance units. a–k,m, Scale bars, 10 μm. a–d,f,h,i,l,n, Data are representative of three independent experiments with similar results. e,g,j,k,m, Data are presented as the mean ± s.d. j,k,m, Statistical analyses were performed using two-tailed Student’s t-test. DIC, differential interference contrast microscopy; DAPI, 4,6-diamidino-2-phenylindole.
To evaluate whether SARS2-NP undergoes LLPS in cells, we transfected HeLa cells with GFP–SARS2-NP and observed that GFP–SARS2-NP formed discrete puncta in the cytoplasm (Fig. 1f,g). In line with the in vitro results, the GFP–SARS2-NP-enriched droplets in cells readily underwent fusion (Fig. 1f) and recovered quickly following FRAP (Fig. 1g). Because RNA is required for viral ribonucleoprotein assembly, we evaluated whether RNA is involved in GFP–SARS2-NP droplet formation by co-transfecting Cy5-labelled SARS-CoV-2 open reading frame (ORF) RNA with GFP–SARS2-NP. The GFP–SARS2-NP-enriched puncta were found to co-localize with Cy5–RNA fluorescence (Fig. 1h), and fusion of GFP–SARS2-NP and Cy3–RNA droplets was observed (Fig. 1i).
To determine which domain is required for NP LLPS, a series of truncated SARS2-NPs labelled with GFP were bacterially expressed and purified (Extended Data Fig. 1b). Although deletion of either the IDR or RNA-binding domain barely affected SARS2-NP-droplet formation, the DD-depleted mutant (d_DD) exhibited no or very low LLPS (Fig. 1j). In HeLa cells, GFP-labelled SARS2-NP d_DD was evenly distributed in the cytoplasm, similar to the localization of wild-type (WT) GFP–SARS2-NP after Hex treatment (Fig. 1k). Co-immunoprecipitation revealed that DD deletion prevented SARS2-NP self-association (Fig. 1l). Semi-denaturing detergent agarose gel electrophoresis (SDD–AGE) analysis of the cytosolic fraction revealed a smear of sodium dodecyl sulfate (SDS)-resistant high-molecular-weight SARS2-NP, but not d_DD (Extended Data Fig. 1f,g), in the absence but not in the presence of β-mercaptoethanol (Extended Data Fig. 1f). In sucrose gradients, SARS2-NP WT, but not d_DD, sedimented as a much larger assembly than predicted by its molecular weight (Extended Data Fig. 1h). A proximity ligation assay (PLA)23,24 showed that the Flag– and Myc–SARS2-NP together yielded PLA signals in the cytoplasm, which were detected for the DD-deficient mutant (Fig. 1m). Thus, the DD is required for SARS2-NP condensation in vitro and in vivo.
The NP is highly basic (pI of about 10) and contains multiple RNA-binding sites25,26. Its DD can form a tightly linked dimer with a large RNA-binding groove17,25,27. To evaluate the binding affinity between SARS2-NP and viral RNA, we employed surface plasmon resonance (SPR), where the RNA–SARS2-NP binding affinity was analysed by flowing recombinant SARS2-NP over the RNA-captured sensor chip. As expected, the SARS2-NP exhibited a high binding capacity for viral RNA, which was substantially reduced in the DD-depleted mutant (Fig. 1n), indicating that the DD is required for SARS2-NP to bind viral RNA.
NP-induced innate immune evasion relies on the DD
To explore the function of NP in the innate antiviral response, constructs encoding SARS-NP or SARS2-NP along with IFN-β and PRD I–III (contains the IRF3-binding site of the Ifnb1 promoter) reporter plasmids were transfected into HEK293T cells, followed by Sendai virus (SeV) stimulation. Exogenous expression of both NPs resulted in significant downregulation of IFN-β and PRD I–III promoter activity (Fig. 2a). Overexpression of SARS-NP and SARS2-NP also resulted in reduced expression of IFNB1 messenger RNA and downstream ISGs following induction by SeV (Fig. 2b). Consistent with these results, transfection with both NPs inhibited the 5′-triphosphorylated RNA (5′-ppp RNA)- or poly(I:C)-induced expression of IFNB1 mRNA (Fig. 2c). Furthermore, a slightly stronger inhibitory effect on IFN-β signalling was exhibited by SARS2-NP than by SARS-NP (Fig. 2a–c).
a, Fold change in IFN-β– and PRD I–III–luciferase (luc) activity in HEK293T cells transfected with control vector, and SARS-NP and SARS2-NP expression plasmids, followed by SeV stimulation for 12 h (left). Immunoblot analysis of the expression of SARS-NP and SARS2-NP plasmids in HEK293T cells (right). b, Normalized IFNB1, CCL5 and CXCL10 mRNA expression (determined by qPCR analysis) in HEK293T cells transfected with control vector, or SARS-NP or SARS2-NP expression plasmids, followed by SeV infection for the indicated times. c, Normalized IFNB1 mRNA expression (determined by qPCR analysis) in HEK293T cells transfected with the indicated plasmids and stimulated with 5′-ppp (left) or poly(I:C) (right) for 12 h. d, Normalized IFNB1 mRNA expression (determined by qPCR analysis) in HEK293T transfected with SARS2-NP WT or its truncated plasmids and infected with SeV for 12 h. e, Normalized IFNB1 mRNA expression (determined by qPCR analysis) in HEK293T transfected with the indicated plasmids and stimulated with 5′-ppp RNA (left) or poly(I:C) (right) for various time periods. f, ELISA determination of the IFN-β levels in RAW264.7 (left) and A549 (right) cells transfected with the indicated plasmids and infected with SeV for various time periods (left), and SeV or VSV for 12 h (right). g, Fold change in Ifnb1 mRNA (left) and VSV load (middle; determined by qPCR analysis) as well as the virus titres of VSV (right; determined by plaque assay) in RAW264.7 cells transfected with control, SARS2-NP WT or SARS2-NP d_DD plasmids and infected with VSV (m.o.i. of 0.1) for various time periods. h, Immunoblot of VSV-G in MEF cells transfected with control, SARS2-NP WT or SARS2-NP d_DD plasmids and infected with VSV (m.o.i. of 0.1) for the indicated times (top). The VSV-G band intensities are shown (bottom). i, Bright-field (top) and fluorescence microscopy (bottom) of VSV–GFP in HEK293T cells transfected with control, SARS2-NP WT or SARS2-NP d_DD plasmids, followed by infection with VSV–GFP (m.o.i. of 0.1) for 12 h (left). Scale bar, 100 μm. The fold change in VSV–GFP intensity was quantified using ImageJ (right). j, A549 cells were transfected with the indicated plasmids and infected with SeV for various time periods (top). The rates of SeV-induced cytopathic effect shown were quantified using ImageJ (bottom). a–j, Data are presented as the mean ± s.d.; n = 3 independent samples. Statistical analyses were performed using a two-tailed Student’s t-test (a–i) or two-way analysis of variance (ANOVA; j); NS, not significant.
Deletion of the SARS2-NP DD almost completely abolished the inhibitory effect of SARS2-NP on the expression of IFNB1, CCL5 and CXCL10 mRNA as well as on IFN-β promoter activity (Fig. 2d and Extended Data Fig. 2a,b). The same results were obtained using human lung carcinoma A549 cells (Extended Data Fig. 2c). In line with these findings, overexpression of SARS2-NP d_DD had no obvious effect on IFNB1 mRNA expression induced by poly(I:C) and 5′-ppp RNA (Fig. 2e and Extended Data Fig. 2d). Enzyme-linked immunosorbent assay (ELISA) analysis indicated that the ectopic expression of SARS2-NP WT in RAW264.7 and A549 cells significantly repressed SeV-induced IFN-β secretion, whereas the ectopic expression of SARS2-NP d_DD had no effect (Fig. 2f). After challenge with vesicular stomatitis virus (VSV), an RNA virus, Ifnb1 expression was inhibited and the VSV-specific mRNA, protein and titres (determined as plaque-forming units, p.f.u.) were increased by WT NP but not by NP d_DD (Fig. 2g,h and Extended Data Fig. 2e,f). Accordingly, the replication of VSV expressing GFP (GFP–VSV) was increased by the ectopic expression of WT NP, whereas NP d_DD had no effect on viral replication (Fig. 2i). In addition, ectopic expression of WT NP, but not NP d_DD, maximized the cytopathic effect induced by SeV infection (Fig. 2j and Extended Data Fig. 2g). Thus, SARS2-NP negatively regulates IFN-β signalling and DD is critical for these inhibitory effects.
The DD mediates NP-induced innate immune evasion in vivo
To further elucidate the role of SARS2-NP in the regulation of innate antiviral immunity in mice, we genetically engineered recombinant VSVs by employing the virulence-attenuated VSV backbone harbouring an M51R substitution in the matrix (M) protein28,29. We inserted the coding sequence of control GFP, SARS2-NP WT or SARS2-NP d_DD into the VSV backbone between the VSV glycoprotein (G protein) and the polymerase protein (L protein)30, hereafter denoted as VSV, NP–VSV and NP d_DD–VSV, respectively. This enabled us to perform viral infections in mice systematically and observe the effects of NPs on virulence and antiviral immunity (Fig. 3a). All three viruses displayed similar replication kinetics in plaque assays in Vero E6 cells. However, the NP–VSV replicated more efficiently than the control VSV and NP d_DD–VSV in HEK293T cells (Fig. 3b). Because Vero E6 cells lack an IFN response, the observation described above implied that NP-mediated innate immunosuppression probably contributes to viral proliferation.
a, Schematic of recombinant VSV carrying GFP (as a control), SARS2-NP WT or SARS2-NP d_DD. b, Virus titres (determined by plaque assay) of Vero E6 (left) and HEK293T (right) cells infected with VSV, NP–VSV or NP d_DD–VSV (m.o.i. of 0.1) for various time periods. c, Normalized Ifnb1 mRNA expression (left) and VSV copy number (middle; determined by qPCR analysis) as well as the virus titres of VSV (right; determined by plaque assay) in peritoneal macrophages infected with VSV, NP–VSV or NP d_DD–VSV (m.o.i. of 0.1). b,c, n = 3 biological independent samples. d, Normalized Ifnb1 mRNA expression in the spleen (left), liver (middle) and lungs (right) of mice (n = 5 mice per group) given intraperitoneal (i.p.) injections of VSV, NP–VSV or NP d_DD–VSV (5 × 108 p.f.u. per mouse). e, ELISA determination of the levels of IFN-β in the serum of the mice infected in d. f, Normalized VSV RNA (determined by qPCR analysis) in the spleen, liver and lungs of the mice infected in d. g, Immunoblot analysis of VSV-G in the spleen (top), liver (middle) and lungs (bottom) of the mice infected in d; n = 3 mice. h, Viral titres (determined by plaque assay) of VSV in the spleen, liver and lungs of the infected mice in d. i, Analysis of tissue damage in H&E-stained lung sections from mice treated with PBS or VSV as in d. Scale bars, 500 µm (top) and 100 µm (bottom). Images represent five independent samples with a similar pattern. j, Survival of mice (n = 10 mice per group) at various times after infection with VSV, NP–VSV or NP d_DD–VSV (2 × 109 p.f.u. per mouse; i.p. injection). b–f,h, Data are presented as the mean ± s.d. b–f,h,j, Statistical analyses were performed using a two-tailed Student’s t-test (c–f,h), two-way ANOVA (b) or log-rank test (j); NS, not significant.
We first infected primary peritoneal macrophages with recombinant VSVs at a multiplicity of infection (m.o.i.) of 0.1. After stimulation, Ifnb1 expression was inhibited and the VSV-specific mRNA and VSV titres were increased in cells infected with NP–VSV but not in those infected with NP d_DD–VSV (Fig. 3c). To investigate the function of NP in vivo, mice were challenged with the recombinant VSVs. The levels of Ifnb1 mRNA were significantly lower in the spleen, liver and lungs of the NP–VSV-infected mice than in those of the mice infected with control VSV or NP d_DD–VSV (Fig. 3d). The serum concentration of IFN-β was also lower in NP–VSV-infected mice than control VSV- and NP d_DD–VSV-infected mice (Fig. 3e). In contrast, the VSV-specific mRNA and protein expression as well as the VSV titres were all higher in the lungs, liver and spleen of the NP–VSV-infected mice than the control VSV- and NP d_DD–VSV-infected mice (Fig. 3f–h). Haematoxylin and eosin (H&E) staining of the lungs after infection revealed higher immune-cell infiltration as well as severe tissue injury in the NP–VSV-infected mice than in their control VSV- and NP d_DD–VSV-infected counterparts (Fig. 3i). Moreover, the mice that were infected with recombinant VSV harbouring SARS2-NP WT exhibited substantially higher sensitivity to the virus relative to the mice that were challenged with control VSV. However, such a difference was not observed in the mice that were infected with NP d_DD–VSV (Fig. 3j). These findings indicate that the DD mediates the NP-induced suppression of innate antiviral immunity and contributes to viral replication in vivo.
SARS2-NP negatively regulates MAVS–IRF3 activation and is enriched in MAVS droplets
Next, we investigated whether NP inhibited IFN-β signalling by evaluating virus-induced phosphorylation of TBK1 and IRF3 as well as IRF3 dimerization and its nuclear accumulation31,32. Surprisingly, all of these molecular events were inhibited by ectopic expression of SARS2-NP WT but not SARS2-NP d_DD (Fig. 4a, left). Similar results were found when we analysed SeV-induced signalling events in A549 cells (Fig. 4a, right). Moreover, cells expressing SARS2-NP WT, but not d_DD, exhibited severely impaired IRF3 nuclear translocation after stimulation with SeV (Fig. 4b,c). Thus, SARS2-NP might target critical IFN-β signalling components upstream of TBK1–IRF3.
a, Immunoblot of IRF3 dimerization (top; native gel) and p-IRF3, p-TBK1, p-IKKε and total IRF3, TBK1 and IKKε in HEK293T (left) and A549 cells (right) transfected with control, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids and infected with SeV for various time periods. b, Immunofluorescence microscopy and DAPI staining of U2OS cells transfected with the control, SARS2-NP WT or SARS2-NP d_DD plasmids, followed by infection with SeV for 8 h. c, Immunoblot analysis of the nuclear and cytoplasmic fractions of HEK293T (left) and A549 (right) cells transfected with control, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids and infected with SeV for various time periods. Cytoplasmic actin and nuclear laminin B (lamin B) serve as controls. Nuc, nuclear; cyt, cytoplasmic. d, Immunoblot of the TCL and immunoprecipitates derived from HEK293T cells transfected with Myc–SARS2-NP along with Flag-tagged RIG-I, MAVS, TRAF3, TBK1, IKKε and IRF3. e, Immunoblot of the TCL and immunoprecipitates of control IgG or anti-NP antibodies derived from Caco-2 cells infected with SARS-CoV-2. f, Immunoblot of the TCL and immunoprecipitates derived from HEK293T cells transfected with Myc–SARS2-NP along with Flag–MAVS WT or truncated plasmids (depleted CARD (d_CARD), Pro-rich (d_Pro) and transmembrane (d_TM) domains or a combination thereof (d_CARD + Pro + TM and d_CARD + Pro)). g, Immunoblot of the TCL and anti-IgG (ns) or anti-MAVS immunoprecipitants derived from HEK293T (left) and A549 cells (right) transfected with control, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids and infected with SeV for 6 h. h, GFP–MAVS (green) and mCherry–NP (magenta; 200 nM each) were mixed with the indicated module concentration and imaged for fluorescence. b,h, Scale bars, 10 μm. a–h, Images are representative of three independent experiments. IB, immunoblot; IP, immunoprecipitation.
To determine which signalling protein could be targeted by SARS2-NP, expression plasmids encoding a series of essential antiviral signalling proteins were co-expressed with SARS2-NP. We found that ectopically expressed SARS2-NP specifically interacted with MAVS (Fig. 4d). Endogenous MAVS immunoprecipitated together with SARS2-NP in cells infected with SARS-CoV-2 (Fig. 4e). MAVS is comprised of a CARD (amino acids 1–90), a Pro-rich (amino acids 91–172) and a transmembrane (amino acids 513–540) domain (Extended Data Fig. 3a). To determine which MAVS domain mediates its association with SARS2-NP, a series of truncated MAVS variants were constructed for interaction assays with SARS2-NP. Deletion of the critical N terminus of MAVS—removal of either the CARD domain or the Pro-rich region—impeded the MAVS–SARS2-NP interaction (Fig. 4f). Following infection with RNA virus, MAVS serves as a central antiviral signalling hub via Lys63-linked poly-ubiquitination by the E3 ligase TRIM31 and aggregates to stimulate downstream signalling cascades. Interestingly, the CARD domain of RIG-I interacts with the CARD domains of MAVS33, whereas TRIM31 interacts with MAVS through the Pro-rich domain of MAVS34. We therefore tested whether SARS2-NP regulates the interaction of MAVS with RIG-I and TRIM31. We indeed observed endogenous TRIM31 and RIG-I associated with endogenous MAVS after infection with SeV in both HEK293T and A549 cells. Furthermore, the infection-induced associations were impeded by SARS2-NP WT, whereas SARS2-NP d_DD had no effect (Fig. 4g). Thus, SARS2-NP suppresses innate immune signalling at the level of MAVS.
MAVS was reported to form clusters in vivo and prion-like aggregates in vitro33; this also suggests that MAVS might undergo LLPS and thus benefit antiviral signalling transduction33,35. In cells, MAVS clusters were found to be disrupted into segments by SARS2-NP WT, but not by SARS2-NP d_DD, and those segments could still partially co-condensate with SARS2-NP WT (Extended Data Fig. 3b). Purified GFP–MAVS formed microsized droplets spontaneously in vitro, and the droplets were larger at higher protein concentrations (Fig. 4h). We next evaluated the potential relationship between SARS2-NP LLPS and its inhibitory regulation of MAVS. Notably, when the MAVS protein was present in excess (above the diagonal), GFP–MAVS was enriched in the droplets (Fig. 4h). In contrast, when SARS2-NP was in excess (below the diagonal), the MAVS–MAVS enrichment pattern was displaced by MAVS–SARS2-NP and SARS2-NP–SARS2-NP enrichment patterns, in which mCherry–SARS2-NP exhibited an opposite pattern of enrichment (Fig. 4h). In contrast, SARS2-NP d_DD barely affected the MAVS droplet formation (Extended Data Fig. 3c). Thus, SARS2-NP negatively regulates MAVS–IRF3 signalling cascades by binding MAVS and interfering with its phase separation.
SARS2-NP inhibits Lys63-linked poly-ubiquitination and aggregation of MAVS
Due to the inhibitory effect of SARS2-NP on the association between MAVS and TRIM31, we observed that ectopic SARS2-NP expression repressed MAVS poly-ubiquitination with WT and Lys63-linked ubiquitin (Fig. 5a). Using tandem ubiquitin-binding entities (TUBE), we pulled-down endogenous K63-ubiquitin-conjugated MAVS from SeV-stimulated HEK293T and A549 cells, which was severely impeded by SARS2-NP WT but not by SARS2-NP d_DD (Fig. 5b,c).
a, Immunoblot of the TCL and immunoprecipitates derived from HEK293T cells transfected with WT, K48- or K63-linked haemagglutinin (HA)–ubiquitin (Ub) plasmids along with Flag–MAVS, TRIM31, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids, as indicated. b,c, Immunoblot of the TCL and TUBE1 pull-down products from HEK293T (b) and A549 (c) cells transfected with control, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids, followed by infection with or without SeV for 12 h. d, SDD–AGE analysis of MAVS aggregation (top) and SDS–PAGE (bottom) of the TCL derived from HEK293T cells transfected with plasmids expressing Flag–MAVS and Myc–SARS2-NP WT or Myc–SARS2-NP d_DD. e,f, SDD–AGE analysis of MAVS aggregation (top) and SDS–PAGE (below) of TCL derived from HEK293T (e) and A549 cells (f) transfected with control, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids, followed by infection with SeV for different time periods (e) or 12 h (f). g, Sucrose-gradient ultracentrifugation analysis of crude mitochondria isolated from HEK293T cells transfected with control, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids and infected with or without SeV for 12 h (top). The MAVS band intensities are shown (bottom; mean ± s.d.). h, Immunoblot of the TCL and immunoprecipitates derived from HEK293T cells transfected with expression plasmids for Flag–MAVS, SARS2-NP WT or SARS2-NP d_DD along with Myc–TBK1 (left) or Myc–IKKε (right). i, Immunoblot of the TCL and anti-IgG (ns) or anti-MAVS immunoprecipitates derived from HEK293T (left) and A549 cells (right) transfected with control, Myc–SARS2-NP WT or Myc–SARS2-NP d_DD plasmids and infected with SeV for 6 h. a–i, Data are representative of three independent experiments. IB, immunoblot; IP, immunoprecipitation.
MAVS aggregation relies on Lys63-linked poly-ubiquitination induced by virus infection that also recruits TBK1 and IKKε for phosphorylation and activation34,36. Consistent with impaired ubiquitination of MAVS, we found that the aggregation of MAVS—either transfected or at endogenous levels induced by SeV—can be suppressed by the ectopic expression of SARS2-NP WT but not SARS2-NP d_DD (Fig. 5d–f). In sucrose-gradient ultracentrifugation, SeV infection strongly shifted MAVS from the low-molecular-weight fractions to the high-molecular-weight fractions, and this shift was severely impaired in cells with ectopic SARS2-NP WT expression but not in those with ectopic expression of SARS2-NP d_DD (Fig. 5g). Consistent with these results, SARS2-NP WT, but not SARS2-NP d_DD, strongly interfered with the association between co-expressed MAVS and TBK1/IKKε (Fig. 5h). At endogenous levels, SeV-induced MAVS–TBK1–IKKε complex formation in both HEK293T and A549 cells was disrupted by SARS2-NP WT but not by SARS2-NP d_DD (Fig. 5i). Together, these results indicate that SARS2-NP strongly inhibits MAVS activation in cells.
SARS2-NP acetylation at Lys375 abrogates its LLPS and the NP-mediated suppression of MAVS signalling
These findings raised the question of how hosts manage to mount a defence against viral infections in the presence of high amounts of NP. We therefore investigated whether NP itself is regulated in the host. In fact, we noticed that a few mice infected with NP–VSV seemed to have mild pathological symptoms compared with others and exhibited only slight weight loss. Thus, we extracted NP from the tissue of these mice for mass spectrometry analysis. We observed that SARS2-NP can be highly acetylated in vivo at Lys375 (Fig. 6a), a motif that is adjacent to the DD and conserved between SARS-CoV-NP and Middle East respiratory syndrome coronavirus (MERS-CoV)-NP (Fig. 6b). Using an acetyl-Lys antibody, we confirmed that abundant NP can be acetylated in host cells, and Lys375 was the most critical residue for NP acetylation (Fig. 6c). Co-expression of CREB-binding protein (CBP), but not the other acetyltransferases tested, catalysed NP acetylation at Lys375 (Extended Data Fig. 4a,b). Furthermore, knockdown of endogenous CBP with independent small interfering RNA (siRNA) significantly inhibited NP acetylation (Extended Data Fig. 4c), indicating that host CBP mediates NP acetylation at Lys375. To consolidate that NP was acetylated at Lys375, we generated an antibody specific to Lys375-acetylated NP and found that cells treated with either the sirtuin inhibitor nicotinamide (NAM) or the HDAC inhibitor trichostatin A showed increased Lys375 acetylation of NP (Extended Data Fig. 4d), revealing that both sirtuins and HDACs might be involved in deacetylating the NP. Among the eleven HDACs (HDAC1–11) and seven sirtuins (SIRT1–7) tested, multiple enzymes—including HDAC6, -7 and -9, and Sirt1, -3, -4 and -5—were found to effectively deacetylate NP (Extended Data Fig. 4e).
a, SARS2-NP was immunoprecipitated from NP–VSV-infected mouse tissues with antibody to SARS2-NP (left). Mass spectrometry analysis of a representative SARS2-NP peptide carrying acetylated Lys375 (right). b, Diagram of the SARS2-NP acetylation sites (top) and sequence alignment of the Lys375 in NP orthologues from different coronaviruses (bottom). c, Immunoblot of control IgG or anti-acetyl-Lys (Acetyl-K) immunoprecipitates derived from HEK293T cells transfected with the indicated expression plasmids. d, Streptavidin RNA pull-down of Myc–SARS2-NP WT, K375Q and K375R (left). The fold changes in RNA-bound Myc–SARS2-NP are shown (right). e, In situ PLA of Myc– and Flag–SARS2-NP in HeLa cells transfected with the indicated plasmids and infected with SeV for 8 h (left). The PLA-detected proximity (PROX) complexes (red dots) were quantified (right). f, Immunoblot analysis of TUBE1 pull-down products (left) and SDD–AGE analysis of MAVS aggregation (right) in A549 cells transfected with the indicated plasmids and infected with SeV for 12 h. g, Normalized IFNB1 mRNA expression (determined by qPCR analysis) in A549 cells transfected with the indicated plasmids and infected with SeV or VSV for 12 h (n = 3 biologically independent samples). h, Schematic describing the generation of site-specific acetylated recombinant SARS2-NP with an acetyl-Lys tRNA (left). Immunoblot of the purified protein with antibody specific to acetyl-Lys375 of SARS2-NP (right). i, Binding affinity, determined using an SPR assay, between RNA and non-acetylated SARS2-NP (left) or SARS2-NPLys375-Ac (right) purified from h; r.u., resonance units. j, Droplet formation of AF488-labelled SARS2-NP WT (non-Ac) or SARS2-NPLys375-Ac (5 µM) mixed with Cy3–RNA (5 µM). Fold change in droplet formation (right; n = 6 independent experiments). DIC, differential interference contrast microscopy. k, FRAP quantification of SARS2-NP non-Ac or Lys375-Ac (5 µM) and RNA (5 µM) droplets before and after photobleaching over 35 s (n = 3 droplets). e,j, Scale bars, 10 μm. a,c–f,h,i, Data are representative of three independent experiments with similar results. d,e,g,j,k, Data are presented as the mean ± s.d., Statistical analyses were performed using a two-tailed Student’s t-test (d,e,g,j) or two-way ANOVA (k). IB, immunoblot; IP, immunoprecipitation.
We next generated a mutant by replacing Lys with Gln, thus mimicking the acetylation state of Lys. Although SARS2-NP WT exhibited efficiency in the RNA-binding assay, a K375Q substitution strongly inhibited NP activity (Fig. 6d). Moreover, a K375R replacement resulted in a greater than fivefold increase in RNA-binding activity compared with the WT control (Fig. 6d). In line with its role in regulating Lys375 acetylation, gain or loss of CBP expression significantly enhanced or reduced the RNA-binding affinity of SARS2-NP, respectively (Extended Data Fig. 4f,g). Compared with SARS2-NP WT, the K375Q mutant exhibited severely reduced PLA signals both at the basal and virus-induced levels (Fig. 6e). Moreover, the K375Q mutant partially lost activity towards Lys63-linked poly-ubiquitination and aggregation of MAVS, whereas K375R exhibited enhanced ability (Fig. 6f). Consistent with this, the K375Q mutant was less efficient in suppressing IFN-β promoter activity and IFNB1 mRNA production as well as in enhancing GFP–VSV replication than SARS2-NP WT, whereas these activities were significantly enhanced in the K375R mutant (Fig. 6g and Extended Data Fig. 4h,i). Therefore, the NP-mediated attenuation of MAVS activity and antiviral signalling could be partially rescued by Lys375 acetylation.
To verify the effect of acetylation on NP function, we employed genetic code expansion orthogonal systems37,38 to incorporate acetyl-Lys into a recombinant SARS2-NP to create a fully Lys375-acetylated NP (SARS2-NPLys375-Ac). In this system, a pBAD/Myc–His vector carrying the ORF for SARS2-NP with an amber codon at the desired site for Lys375 was co-transformed with a plasmid harbouring aminoacyl-transfer RNA synthetase (pSupAR-tRNA-chRS42) into the Escherichia coli strain DH10B. The E. coli were subsequently treated with NAM and Nε-acetyl-Lys and induced with l-arabinose to express fully Nε-acetylated Lys375 NP (SARS2-NPLys375-Ac; Fig. 6h, left); SARS2-NP WT in the same vector were purified as non-acetylated control (SARS2-NPNon-Ac; Fig. 6h, left).
As expected, antibodies specific to Lys375-acetylated NP can detect SARS2-NPLys375-Ac but not SARS2-NPNon-Ac (Fig. 6h, right). In SPR assays, SARS2-NPLys375-Ac was much less efficient than SARS2-NPNon-Ac in binding to viral RNA in vitro (Fig. 6i), indicating that Lys375-acetylated NP is indeed less active. To test whether acetylation alters phase separation of NP in vitro, we incubated fluorescently labelled NP with viral RNA. SARS2-NPNon-Ac and RNA rapidly formed micrometre-sized liquid droplets and the small liquid droplets quickly fused into larger ones, accompanied by increased fluorescence intensity and a larger equivalent diameter. In contrast, SARS2-NPLys375-Ac and RNA slowly formed smaller droplets with much less liquid (Fig. 6j). FRAP experiments showed that following bleaching, SARS2-NPNon-Ac binding to RNA was recovered more efficiently than that of SARS2-NPLys375-Ac (Fig. 6k). Thus, NP acetylation at Lys375 abrogates its LLPS.
K375N, K375E and K375Q variants that mimic acetylation exhibit reduced LLPS and ability to suppress innate antiviral immunity
RNA viruses have very high mutation rates, enhancing evolvability and altering their virulence39. On 26 May 2021, 1,096,918 SARS-CoV-2 sequences had already been reported and deposited (https://bigd.big.ac.cn/ncov)40,41. Analysis of the corresponding NPs revealed that several variants, including K375Q, and more frequently reported as K375N and K375E, existed for the SARS2-NP (Fig. 7a). Mutation of Lys to Gln, Asn and particularly Glu reduces its positive charge (Gln and Asn are neutral, and Glu has a negative charge), which to a certain extent mimics acetylation of the corresponding Lys residue (Fig. 7a). Considering that Lys375 is adjacent to the DD, a key region responsible for the NP–RNA association, NP LLPS and the NP-mediated blockage of RIG-I-like-receptor signalling, acetylation and acetyl-mimicking mutations at this site probably affect the electrostatic interactions between the DD and RNA, thereby interfering with SARS2-NP function. Thus, the K375N and K375E variants probably attenuate NP-mediated innate immune suppression, in turn reducing virulence. We sought to experimentally confirm this hypothesis.
a, K375 variants in SARS2-NP sequences (top). Schematic of SARS2-NP mutants and RNA valencies (bottom). b, Binding affinity between RNA and SARS2-NP WT (left), K375N (middle) or K375E (right), determined using an SPR assay; r.u., resonance units. c, Immunoblot analysis (left) and fold change in the RNA-binding affinity (right) of streptavidin RNA pull-down derived from HEK293T cells transfected with the indicated plasmids. d, Representative images (left) and quantification (right; fold change) of phase separation of AF488-labelled SARS2-NP WT, K375N or K375E (5 µM) with Cy3–RNA (5 µM; n = 6 independent experiments). DIC, differential interference contrast microscopy. e, In situ PLA (left) and quantification (right) of Myc– and Flag–SARS2-NP in HeLa cells transfected with the indicated plasmids and then infected with SeV for 8 h. The PLA-detected proximity (PROX) complexes were quantified. d,e, Scale bars, 10 µm. f, Immunoblot of the TUBE1 pull-down products from A549 cells transfected with the indicated SARS2-NP plasmids and then infected with SeV for 12 h. g, SDD–AGE analysis of MAVS aggregation of the TCL of A549 cells transfected with the indicated plasmids and infected with SeV for 12 h. h, Normalized Ifnb1 mRNA expression (determined using qPCR analysis) in the spleen (left), liver (middle) and lungs (right) of mice (n = 5 mice per group) given i.p. injection of NP–VSV, NP–VSV K375N, NP–VSV K375E or NP–VSV K375Q (5 × 108 p.f.u. per mouse). i, ELISA determination of the IFN-β levels in the serum of the mice in h. j, Normalized VSV RNA levels (determined using qPCR analysis) in the spleen, liver and lungs of the mice in h. k, Plaque assay of VSV in the spleen, liver and lungs of the mice in h. l, Survival rates of the mice (n = 10 mice per group) infected with NP–VSV WT, K375N, K375E or K375Q (1 × 109 p.f.u. per mouse). m, Normalized IFNB1 mRNA (left) and SARS-CoV-2 genomic RNA (right) levels (determined using qPCR analysis) in Caco-2 cells transfected with the indicated plasmids and then infected with SARS-CoV-2 for 12 h. EV, empty vector. b,c,e–g,m, Data are representative of three independent experiments with similar results. c–e,h–k,m, Data are presented as the mean ± s.d. c–e,h–k,l,m, Statistical analyses were performed using a two-tailed Student’s t-test (c–e,h–k,m) or two-way ANOVA (l). IB, immunoblot.
As expected, the purified K375N and K375E mutants seemed to exhibit reduced binding affinity for viral RNA in SPR and oligo pull-down experiments in vitro, during which a more obvious reduction was observed for K375E (Fig. 7b,c). This was consistent with the idea that K-to-E mutations resulted in a more dramatic charge reduction. To test whether these viral mutations induce changes in the phase separation of SARS2-NP, we incubated fluorescently labelled K375N and K375E with viral RNA and compared the micrometre-sized liquid droplet formation with that of SARS2-NP WT. Both the K375N and K375E mutants seemed to have compromised LLPS (Fig. 7d). All of the Lys375 variants of SARS2-NP—that is, K375N, K375E and K375Q—were less effective in suppressing MAVS droplet formation when they were incubated in vitro (Extended Data Fig. 5).
In cells, PLA revealed much weaker signals for K375N and K375E when compared with WT SARS2-NP (Fig. 7e), which is indicative of the impaired dimerization and self-association of SARS2-NP. An SDD–AGE assay confirmed that the K375N and K375E variants were also less efficient at forming detergent-resistant condensates (Extended Data Fig. 6a). In line with this, both variants partially lost their ability to inhibit the SeV-induced Lys63-linked poly-ubiquitination and aggregation of MAVS (Fig. 7f,g). Furthermore, a significantly reduced ability to suppress IFN-β promoter activity (Extended Data Fig. 6b) and IFNB1 mRNA expression (Extended Data Fig. 6c) was observed for the mutants in addition to a reduced ability to upregulate VSV–GFP replication (Extended Data Fig. 6d). To evaluate the functional consequence of SARS2-NP mutations in vivo, we engineered recombinant VSVs expressing the K375N, K375E and K375Q variants (hereafter, NP–VSV K375N, NP–VSV K375E and NP–VSV K375Q, respectively). Compared with mice infected with NP–VSV, expression of Ifnb1 mRNA was significantly higher in the spleen, liver, and lungs of the mice infected with NP–VSV K375N, NP–VSV K375E and NP–VSV K375Q (Fig. 7h). The concentration of IFN-β in the sera of the mice infected with NP–VSV K375N, NP–VSV K375E or NP–VSV K375Q was also higher (Fig. 7i). The levels of VSV-specific mRNA and the VSV titres were lower in the spleen, liver and lungs of the mice infected with the K375N, K375E and K375Q NP–VSV variants than the NP–VSV-infected mice (Fig. 7j,k). Mice that were infected with the three mutant VSVs had lower mortality rates than the NP–VSV-infected mice (Fig. 7l).
To further determine the role of SARS2-NP mutations at Lys375 in SARS-CoV-2 infection, Caco-2 cells were transfected to express SARS2-NP WT, K375N, K375E or K375Q and subsequently challenged with SARS-CoV-2. As expected, exogenous expression of SARS2-NP WT sharply impeded the expression of IFNB1 mRNA and enhanced the replication of SARS-CoV-2 genomic RNA, whereas the Lys375 mutants poorly inhibited IFN-β production, resulting in a lower SARS-CoV-2 genomic RNA load (Fig. 7m). Together, modifications and mutations of the critical Lys375 residue of SARS2-NP could suppress its function, enhancing the innate antiviral response of the host.
Interfering peptide NIP-V targeting the DD disrupts SARS2-NP LLPS and thus enhances the innate antiviral response both in vitro and in vivo
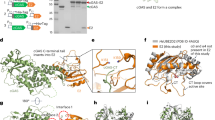
Our findings revealed the indispensable role of the DD of SARS2-NP for SARS2-NP function and immunosuppression. This prompted us to develop a strategy to block the DD. We noticed that protein domains containing natural l-peptides can be mimicked using d-amino acids with a retro-reversed sequence42. Peptide modified with such d-retroinverso (DRI) isoforms can yield new chemical properties, potentially enhancing the peptide potency43,44,45. Several DRI-modified peptides were well-tolerated and therapeutically effective in a double-blinded, randomized, placebo-controlled phase IIb trial46,47,48 and a phase I trial for the systemic treatment of solid tumours49. This existing evidence provided the rationale for designing a well-tolerated and therapeutically effective anti-SARS2-NP peptide with a DRI conformation. Furthermore, recent reports determined the structure of the SARS2-NP DD dimer17,19, describing that the predominant dimerization interface of each DD subunit included two β strands and η1, α3, α4 and α5 helices, located in five regions of the DD. To prevent SARS2-NP-mediated innate immune suppression, we designed and synthesised multiple interfering peptides with a DRI conformation, named NP interfering peptide (NIP) I–V, all of which were designed to inhibit SARS2-NP dimer formation by blocking different DD interaction regions (Fig. 8a,b). To facilitate the cellular uptake of NIPs, peptides were designed to fuse with HIV-TAT (Fig. 8a), a basic and hydrophilic sequence that allows for the energy-independent cellular uptake of cargo through transient pore formation43,50.
a, NIP-I–V designed to block the DD of SARS2-NP. Red, HIV-TAT sequence. b, Structure of the SARS2-NP DD dimer (PDB code: 7C22). c, Droplet formation of GFP–SARS2-NP (10 μM) mixed with NIPs (5 µM). DIC, differential interference contrast microscopy. d, GFP–SARS2-NP condensates in HeLa cells treated with PBS (top) or NIPs (50 µM; bottom) for 12 h. e, Normalized Ifnb1 (left) and Isg56 (right) mRNA, and VSV RNA (middle) levels— determined by qPCR analysis—in peritoneal macrophages pretreated with NIPs (50 µM) for 2 h and infected with NP–VSV (m.o.i. of 0.1) for 12 h. f, SDD–AGE analysis of HEK293T cells transfected with Myc–SARS2-NP and treated with the indicated concentrations of NIP-V for 12 h. g, MAVS aggregation of peritoneal macrophages transfected with Myc–SARS2-NP, pretreated with 50 (+) or 100 µM (++) NIP-V for 2 h and then infected with SeV for 12 h. h, Normalized IFNB1 mRNA and SARS-CoV-2 RNA levels (subgRNA E, subgenomic RNA4-encoding E gene; determined by qPCR analysis) as well as the SARS-CoV-2 titres in Caco-2 cells pretreated with NIP-V and infected with SARS-CoV-2. i, Normalized Ifnb1 mRNA expression in the spleen (left), liver (middle) and lungs (right; determined by qPCR analysis) of hACE2 transgenic mice (n = 6 (uninfected) or 8 (infected) mice per group) pretreated with PBS or NIP-V (25 mg kg−1; i.p.) for 2 h and given intranasal SARS-CoV-2 inoculation (1 × 105 TCID50 per mouse) for another 16/24 h. j, IFN-β concentration in the serum of the mice in i. k, Log-transformed SARS-CoV2 genomic RNA levels (determined by qPCR) in the mouse tissues from i. l, H&E staining of lung tissues from i. m, Immunohistochemistry assay and H score of the SARS-CoV-2 S protein antigen (brown) of lung sections from i. The nuclei were stained with Mayers haematoxylin (blue). n, SARS2-NP LLPS antagonizes the MAVS-mediated antiviral response, which is rescued by NIP-V. c,d,l,m, Scale bars, 10 µm (c,d), 500 µm (l,m (top)) and 100 µm (l,m (bottom)). c–h,l,m, Data are representative of three (d,f,g) or six (c) independent experiments; n = 3 (e,h) or 6 (l,m) independent biological replicates. Data are presented as the mean ± s.d. e,h–k,m, Statistical analysis were performed using a two-tailed Student’s t-test.
In a phase separation assay, NIP-V was screened out to disrupt droplet formation in vitro (Fig. 8c and Extended Data Fig. 7a). NIP-V treatment hampered the formation of NP-enriched liquid-phase droplets in living cells (Fig. 8d and Extended Data Fig. 7b), thereby establishing the inhibitory role of NIP-V on SARS2-NP LLPS in vivo. Compared with the control, cells treated with NIP-V showed no detectable NP self-association (Extended Data Fig. 7c). We subsequently examined whether this interfering peptide could counteract the inhibitory effect of NP on IFN-β signalling. As expected, treatment with NIP-V significantly enhanced the expression of Ifnb1 and Isg56 while reducing the levels of VSV-specific mRNA (Fig. 8e). Higher NIP-V concentrations were found to inhibit SARS2-NP condensation in cells (Fig. 8f) and thus relieve the NP-mediated suppression of MAVS aggregation (Fig. 8g), indicating that this peptide could probably rescue MAVS activity. Quantitative PCR (qPCR) analysis revealed that NP overexpression blocked IFNB1 mRNA expression induced by RIG-IN and MAVS but not by IRF3-5D. This effect could be partially rescued by NIP-V (Extended Data Fig. 7d), confirming the innate immunity-enhancing effect of this peptide.
To determine the function of NIP-V in vivo, mice that had been pre-injected with or without NIP-V were infected with NP–VSV. The expression of Ifnb1 and Isgs mRNA was significantly higher in the spleen, liver and lungs of the NIP-V-treated mice compared with the PBS-treated mice (Extended Data Fig. 8a,b). An ELISA assay revealed that the concentration of IFN-β was also significantly higher in the serum of the NIP-V-treated mice (Extended Data Fig. 8c). Consistent with these results, NIP-V treatment considerably reduced the VSV-specific mRNA levels, VSV-specific protein VSV-G expression and NP–VSV titres in the lungs, liver and spleen of mice (Extended Data Fig. 8d–f). Notably, the mice that were pretreated with NIP-V seemed to be less sensitive to NP–VSV infection than those pretreated with PBS (Extended Data Fig. 8g). Staining with H&E confirmed that there was less damage in the lungs of the NIP-V-treated mice (Extended Data Fig. 8h). Importantly, to determine whether blocking NP could inhibit SARS-CoV-2 replication, we treated SARS-CoV-2-infected CaCo-2 cells with NIP-V. We observed a gradual dose-dependent elevation of IFNB1 mRNA expression following NIP-V treatment, which consequently suppressed active viral replication (as assessed by the levels of the SARS-CoV-2 subgenomic RNA4-encoding E gene) and viral titres (Fig. 8h). To further evaluate the function of NIP-V in the context of SARS-CoV-2 infection, human ACE2 (hACE2) transgenic mice were pre-injected with NIP-V or PBS and then intranasally inoculated with SARS-CoV-2 at 50% tissue culture infectious dose (TCID50) of 1 × 105. Compared with the PBS-treated mice, SARS-CoV-2-induced Ifnb1 and Isg56 mRNA expression in the spleen, liver and lungs were higher in the NIP-V-treated mice (Fig. 8i and Extended Data Fig. 8i). NIP-V treatment also potentiated the IFN-β induction in serum of SARS-CoV-2-infected mice (Fig. 8j). Consistent with these results, the levels of SARS-CoV-2 genomic RNA in the spleen, liver and lungs were drastically reduced in the NIP-V-treated mice (Fig. 8k). Staining with H&E indicated that SARS-CoV-2 infection led to severe lung damage, characterized by thickened alveolar septa and inflammatory-cell infiltration in the PBS-treated hACE2 transgenic mice, whereas only minor lung damage was observed in the mice pretreated with NIP-V (Fig. 8l). Consistent with this, the expression of the S (Fig. 8m) and N (Extended Data Fig. 8j) proteins in the lung were remarkably reduced in the NIP-V-treated mice compared with the PBS-treated mice. Together, the results from the in vitro and in vivo analyses demonstrated that NIP-V could disrupt SARS2-NP LLPS and thus alleviate the NP-mediated suppression of the innate antiviral responses (Fig. 8n).
Discussion
The IFN-I response has been demonstrated to protect the host against SARS-CoV and MERS-CoV in various animal models, including mice, marmosets and macaques51,52,53. The addition of exogenous IFN-I to cell cultures efficiently inhibits the proliferation of SARS-CoV, MERS-CoV and SARS-CoV-2 (refs. 51,54,55,56,57,58,59). To survive in the host, viruses employ various immune evasion mechanisms to counteract innate immune sensing and IFN-I production. Compared with other respiratory RNA viruses, SARS-CoV-2 exhibits an increased immunosuppressive ability and induces low levels IFN-I production both in vitro and in vivo54,60. In the serum of patients with COVID-19, IFN-I expression could not be detected by commonly used assays, despite efficient replication of SARS-CoV-2 in human lung tissue1,54,61,62, which suggests that SARS-CoV-2 harbours a highly efficient mechanism for the inhibition of innate antiviral immunity. However, although several SARS-CoV-2 viral proteins were shown to be implied in immune evasion63, the precise evasion mechanism remains elusive. In this study we demonstrated that the introduction of SARS2-NP, but not SARS2-NP d_DD, leads to a compromised MAVS signalling response, increased viral proliferation and higher mortality in mice. This finding is in agreement with that of a previous report stating that the virulence and clinical severity of coronaviruses are associated with immune evasion64. It is known that human coronavirus 229E induces high levels of IFN and thus exhibits mild virulence65, whereas SARS-CoV and MERS-CoV utilize multiple mechanisms to repress the IFN response, contributing to their high lethality66,67,68.
While our manuscript was in consideration, other groups reported that SARS2-NP undergoes LLPS with RNA26,39,69,70,71,72,73,74, which contributes to virion assembly as well as viral transcription and replication59,60,61. Our findings indicate that SARS2-NP LLPS is associated with its ability to inhibit IFN-I signalling. Notably, we observed SARS2-NP acetylation and deacetylation at Lys375 by host acetyltransferases and deacetylases, which in turn modulated NP self-condensation and NP-regulated MAVS activity. Importantly, we found that the patient-derived SARS-CoV-2 virus was frequently mutated at SARS2-NP Lys375—including the K375N, K375E and K375Q variants, all of which had a reduced positive charge and thus mimicked acetylation. Furthermore, SARS2-NP harbouring these mutations exhibited lower levels of RNA binding, reduced LLPS and reduced ability to suppress innate antiviral immunity.
In this study we designed interfering peptides against all interaction structures within the DD of SARS2-NP. One of the peptides, NIP-V, was screened out as a potent in vitro inhibitor of SARS2-NP LLPS, rescuing MAVS activity and the antiviral immune response in vivo. Thus, the current findings highlight SARS2-NP LLPS as a promising therapeutic target for rescuing innate antiviral immunity during SARS-CoV-2 infection. We believe these insights may help curb the ongoing pandemic as well as any future outbreaks.
Methods
Mice
Mice were maintained under specific-pathogen-free conditions in the animal facility of Soochow University. The hACE2 transgenic mice were purchased from Shanghai Model Organisms Center, Inc. Animal experiments involving infectious virus were performed in the physical containment level 2 laboratory of Soochow University. The Institutional Committee for Animal Welfare of Soochow University approved and oversaw this study. Authentic SARS-CoV-2 challenge studies were approved by the Ethics Committee of ZSSOM of Sun Yat-sen University on Laboratory Animal Care. All of the authentic SARS-CoV-2-related experiments were conducted in a biosafety-level-3 facility. All of the animal studies were conducted in accordance with the Soochow University guidelines or in compliance with the guidelines and regulations of the Laboratory Monitoring Committee of Guangdong Province of China.
Cells and reagents
HEK293T, HeLa, A549, U2OS, Vero E6 and Caco-2 cells obtained from the American Type Culture Collection were cultured at 37 °C under 5% CO2 in DMEM medium supplemented with 10% fetal bovine serum, 100 U ml−1 penicillin and 100 μg ml−1 streptomycin. Peritoneal macrophages were harvested from mice 4 d after injection of thioglycolate (BD) and cultured in DMEM supplemented with 5% fetal bovine serum. All cells were negative for mycoplasma. Poly(I:C) and 5′-ppp RNA were purchased from Invivogen. MG132 was purchased from SelleckChem.
Plasmids and transfection
Expression plasmids for RIG-I, MAVS, TRAF3, TBK1, IKKɛ and IRF3, and plasmids for the IFNB1-Luc and ISRE-Luc transcriptional reporters were described previously75. SARS-CoV NP and SARS2-NP were synthesised by GENEWIZ, China. Sequences encoding the NPs were amplified by standard PCR and cloned into the pcDNA3.1-Myc and pLV-Flag vectors. Point mutations were generated by site-directed mutagenesis with KOD plus polymerase (Toyobo) and the plasmids encoding the WT protein were used as the template. All constructs were confirmed by DNA sequencing. Standard polyethylenimine was used for transient transfection of plasmids and siRNA into HEK293T cells. Plasmids and RNA mimics were transfected into macrophages using Geneporter 2 transfection reagent (Genlantis).
Transcription reporter assay
HEK293T cells were seeded into 48-well plates, transfected with plasmids using polyethylenimine, and subsequently (mock) treated for various times—as indicated in the figures—and harvested for the measurement of luciferase activity. The internal transfection control Renilla was used to normalize the luciferase activities to control for differences in transfection efficiency.
Real-time qPCR with reverse transcription
Total RNA was prepared using the RNAiso Plus kit (Takara). A total of 1 μg RNA was reverse-transcribed using a PrimeScript RT reagent kit (Perfect Real Time; Takara). Real-time PCR was conducted using TB Green premix Ex Taq (Tli RNaseH Plus; Takara) using a StepOne Plus real-time PCR system (Applied Bioscience). Quantitation of all target gene expression was normalized to the control gene Gapdh for mouse genes and GAPDH for human genes.
The qPCR primers were as follows: murine Ifnb1 forward, 5′-TCCTGCTGTGCTTCTCCACCACA-3′; murine Ifnb1 reverse, 5′-AAGTCCGCCCTGTAGGTGAGGTT-3′; murine Cxcl10 forward, 5′-ATCATCCCTGCGAGCCTATCCT-3′; murine Cxcl10 reverse, 5′-GACCTTTTTTGGCTAAACGCTTTC-3′; murine Ccl5 forward, 5′-TCACCATATGGCTCGGACACCAC-3′; murine Ccl5 reverse, 5′-TTGGCACACACTTGGCGGTTC-3′; murine Isg56 forward, 5′-AAGACAAGGCAATCACCCTCTACT-3′; murine Isg56 reverse, 5′-GTCTTTCAGCCACTTTCTCCAAA-3′; murine Gapdh forward, 5′-GGCCTTCCGTGTTCCTACC-3′; murine Gapdh reverse, 5′-AGCCCAAGATGCCCTTCAGT-3′; VSV forward, 5′-ACGGCGTACTTCCAGATGG-3′; VSV reverse, 5′-CTCGGTTCAAGATCCAGGT-3′; human IFNB1 forward, 5′-CCAACAAGTGTCTCCTCCAAAT-3′; human IFNB1 reverse, 5′-AATCTCCTCAGGGATGTCAAAGT-3′; human GAPDH forward, 5′-AGGGCTGCTTTTAACTCTGGT-3′; human GAPDH reverse, 5′-CCCCACTTGATTTTGGAGGGA-3′. human ISG56 forward, 5′-GCTTTCAAATCCCTTCCGCTAT-3′; human ISG56 reverse, 5′-CTTGGCCCGTTCATAATTTTTTC-3′; human CXCL10 forward, 5′-TTTGCTGCCTTATCTTTCTGACT-3′; human CXCL10 reverse, 5′-ATTGTAGCAATGATCTCAACACG-3′; SARS2 E forward, 5′-CTTTCGTGGTATTCTTGCTAGTT-3′ and SARS2 E reverse, 5′-CACGTTAACAATATTGCAGCA-3′.
Immunoprecipitation and immunoblot analyses
Cells were lysed with 1 ml lysis buffer (20 mM Tris–HCl, pH 7.4, 2 mM EDTA, 25 mM NaF and 1% Triton X-100) containing protease inhibitors (Sigma) for 10 min at 4 °C. Immunoprecipitation was performed with anti-Flag M2 beads (Sigma, A2220) for 1 h at 4 °C or with various antibodies (see below) and protein A–sepharose (GE Healthcare Bio-Sciences AB) for 3 h at 4 °C. Thereafter, the precipitates were washed three times with washing buffer (50 mM Tris–HCl, pH 8.0, 150 mM NaCl, 1% Nonidet P-40, 0.5% sodium deoxycholate and 0.1% SDS) and the immunocomplexes were eluted with sample buffer containing 1% SDS for 5 min at 95 °C. Immunoblot analysis was performed with specific antibodies and secondary anti-mouse or anti-rabbit antibodies conjugated to horseradish peroxidase (HRP). Visualization was achieved with chemiluminescence. The antibodies used for immunoprecipitation (IP), immunoblotting (IB) and immunofluorescence (IF) were as follows: mouse monoclonal antibody to SARS2-NP (ABclonal, A20142; 1:2,000 for IB), rabbit monoclonal antibody to MAVS (Cell Signaling, 4983 and 3993; 1:1,000 for IB), anti-acetyl-Lys (Abcam, ab80178; 1:2,000 for IB), MAVS antibody (provided by F. Hou; 1:200 for IP), rabbit monoclonal antibody to IRF3 (Cell Signaling, D614C; 1:1,000 for IB and 1:200 for IF), rabbit monoclonal antibody to phospho-IRF3(Ser 396) (Cell Signaling, 4D4G; 1:1,000 for IB), rabbit monoclonal antibody to TBK1 (Cell Signaling, D1B4; 1:1,000 for IB), rabbit monoclonal antibody to phospho-TBK1(Ser172) (Cell Signaling, D52C2; 1:1,000 for IB), rabbit monoclonal antibody to IKKɛ (Cell Signaling, D20G4; 1:1,000 for IB), rabbit monoclonal anti-phospho-IKKɛ (Ser172) (Cell Signaling, 8766; 1:1,000 for IB), mouse monoclonal anti-Flag (M2) (Sigma-Aldrich, F3165; 1:2,000 for IB and 1:200 for IF), rabbit polyclonal anti-HA (Y11) (Santa Cruz Biotechnology, sc-805; 1:2,000 for IB), rabbit monoclonal anti-HA (12CA5) (made in-house; 1:2,000 for IB), rabbit polyclonal anti-Myc (A-14) (Santa Cruz Biotechnology, sc-789; 1:2,000 for IB), rabbit polyclonal anti-CBP (Santa Cruz Biotechnology, sc-583; 1:1,000 for IB), mouse monoclonal anti-Myc (9E10) (Santa Cruz Biotechnology, sc-40; 1:2,000 for IB), mouse monoclonal anti-actin (Sigma-Aldrich, AC-15; 1:2,000 for IB), anti-VSV-G (ABGENT, AP1016a; 1:1,000 for IB), HRP-mouse anti-biotin (Invitrogen, 03-3720; 1:1,000), protein A–HRP (Sigma-Aldrich, GENA9120; 1:10,000 for IB) and HRP-conjugated secondary antibodies (Cell Signaling, 7076 (anti-mouse IgG) or 7074 (anti-rabbit IgG); 1:10,000 for IB); the polyclonal antibody to acetylated Lys375 of SARS2-NP was generated by immunizing rabbits with Cys-KDKKK(Kac)ADETQAL peptide (1:2,000 for IB and 1:200 for IP). Fluorescein AffiniPure goat anti-mouse IgG (H + L) (115-095-146), rhodamine (TRITC) AffiniPure goat anti-mouse IgG (H + L) (115-025-146), fluorescein AffiniPure goat anti-rabbit IgG (H + L) (111-095-003) and rhodamine (TRITC) AffiniPure goat anti-rabbit IgG (H + L) (111-025-003) were from Jackson ImmunoResearch.
SDD–AGE
The SDD–AGE protocol was previously described36. Briefly, cells were lysed in Buffer A (10 mM Tris–HCl, pH 7.5, 10 mM KCl, 1.5 mM MgCl2, 0.25 M d-mannitol and protease inhibitor cocktail) by repeated douncing, and the homogenized cells were centrifuged at 700g for 10 min at 4 °C. The supernatant was transferred and centrifuged at 10,000g for 30 min at 4 °C to form a pellet, which was the intact crude mitochondrion (P5). The crude mitochondria (P5) were resuspended in 1×sample buffer (0.5×TBE, 10% glycerol, 2% SDS and 0.0025% bromophenol blue) and loaded onto a vertical 1.5% agarose gel. After electrophoresis in the running buffer (1×TBE and 0.1% SDS) for 35 min with a constant voltage of 100 V at 4 °C, the proteins were transferred to an Immobilon membrane (Millipore) for immunoblotting.
Protein fractions in sucrose gradient
Protein-fraction isolation on a sucrose gradient was previously described36. Briefly, crude mitochondria (P5) isolated as described in the ‘SDD–AGE’ section were resuspended in Buffer B (20 mM HEPES (pH 7.4), 10% glycerol, 0.5 mM EGTA and 0.25 M d-mannitol) and loaded on top of a centrifuge tube containing 1 ml of 50% sucrose in PBS on the bottom layer and 1 ml of 40% sucrose in PBS on the top layer. After centrifugation at 100,000g for 30 min, the mitochondria enriched at the interface of the two layers were collected and solubilized in PBS containing 1% n-dodecyl-β-d-maltosid. The mitochondrial lysate was loaded onto a sucrose gradient (10–60%) and centrifuged at 160,000g for 2 h. Seven fractions of equal volume were taken from the top to bottom of the tube and loaded on the SDS–PAGE gel.
TUBE1 assay
The isolation of endogenous K63-linked polyubiquitinated substrates by TUBE1 was previously described76. Equilibrated agarose–TUBEs were incubated with cell lysis buffer for 4 h at 4 °C. The cell lysate was clarified by centrifugation for 10 min at approximately 12,000g (4 °C). The agarose–TUBEs were then washed with TBST and the resin was treated with 0.2 M glycine–HCl, pH 2.5 on a rocking platform for at least one 1 h at 4 °C. The resin was then collected by high-speed centrifugation (13,000g) for 5 min and the supernatant was recovered.
In vivo ubiquitination assay
Cells were washed with PBS and lysed in two pellet volumes of RIPA buffer (20 mM NAP, pH 7.4, 150 mM NaCl, 1% Triton X-100, 0.5% sodium deoxycholate and 1% SDS) supplemented with protease inhibitors and 10 mM N-ethylmaleimide. The lysates were sonicated, boiled at 95 °C for 5 min, diluted with RIPA buffer containing 0.1% SDS and then centrifuged at 4 °C (16,000g for 15 min). The supernatant was incubated with specific antibody and protein A–sepharose for 3 h at 4 °C. After extensive washing, the bound proteins were eluted with 2×SDS sample buffer and separated on an SDS–PAGE gel, followed by western blotting.
Native PAGE
Cells were lysed in RIPA lysis buffer (50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1% NP-40 and 100 μM PMSF) and 15 μg supernatant of the TCL was used for native PAGE after centrifugation; 7.5% native PAGE gels were made without SDS. The gels were pre-run with 25 mM Tris and 192 mM glycine, pH 8.4, with 1% deoxycholate in a cathode chamber for 30 min at 40 mA. Samples in native sample buffer (10 μg protein, 62.5 mM Tris–HCl, pH 6.8, 15% glycerol and 1% deoxycholate) were size-fractionated by electrophoresis for 60 min at 25 mA and transferred to PVDF membranes for immunoblot analysis.
Purification of Lys375-specific fully acetylated SARS2-NP recombinant proteins
The K375 acetylated SARS2-NP recombinant proteins were generated according to a previous report38. After optimizing the constructs and methods, the E. coli DH10B strain was transformed with the plasmid aminoacyl-tRNA synthetase pSupAR-tRNA-chRS42 and pBAD/Myc–His vector carrying the ORF of SARS2-NP with an amber codon at the desired site for Lys375. The cells were first cultured overnight in LB medium supplemented with 100 μg ml−1 ampicillin and 50 mg ml−1 chloramphenicol (Amp+ Cm+) at 37 °C. A 10-ml volume of the overnight bacterial culture was then subcultured in 200 ml LB Amp+ Cm+. When the optical density at 600 nm (OD600) reached 0.6–0.7, 20 mM NAM and 100 mM acetyl-Lys were added, and 30 min later, when the OD600 reached 0.8–1.0, protein expression was induced at 30 °C for 18–20 h by adding 2 mM l-arabinose. The cells were harvested after induction and washed with ice-cold PBS containing 20 mM NAM; the proteins were purified with HisTrap FF (GE Healthcare, 17-5319-01) according to the manufacturer’s protocol.
Immunofluorescence and confocal microscopy
HeLa or U2OS cells cultured on glass coverslips were transfected with plasmids or treated with various stimuli as indicated in the figures. The cells were then washed with PBS, fixed with 4% paraformaldehyde in PBS for 20 min, permeabilized with 0.2% Triton X-100 and blocked with 3% BSA. The cells were then stained with the appropriate antibodies (as listed in the ‘Immunoprecipitation and immunoblot analyses’ section and indicated in the figures), followed by incubation with fluorescent-dye-conjugated secondary antibodies. The nuclei were counterstained with DAPI (Sigma-Aldrich). The tissue paraffin sections were blocked with 3% BSA and 0.1% Tween-20 in PBS for 1 h after rehydration and antigen retrieval, and the sections were then stained with the indicated antibodies in 3% BSA, followed by incubation with fluorescent-dye-conjugated secondary antibodies in 3% BSA. The nuclei were counterstained with DAPI in 3% BSA.
In vitro phase separation assay
Purified proteins were diluted to various concentrations in buffer containing 20 mM Tris–HCl pH 5.5 and 1 mM dithiothreitol with the indicated salt concentrations. Protein solution (5 μl) was loaded onto a glass slide, covered with a coverslip and imaged using an LSM880 confocal microscope system (Zeiss). Phase separation of recombinant EGFP–SARS2-NP with Cy3-labelled RNA was performed in physiological buffer (20 mM Tris–HCl, pH 7.0, 15 mM NaCl, 135 mM KCl, 5 mM phosphate, 1.5 mM MgCl2 and 1 mg ml−1 BSA). The mixture was transferred to 96-well plates (Corning) coated with 20 mg ml−1 BSA (Sigma) and imaged.
The sequence of the Cy3- or Cy5-labelled SARS-CoV-2 RNA was as follows: 5′-CACUCGCUAUGUCGAUAACAACUUCUGUGGCCCUGAUGGCUACCCUCUUGAGUGCAUUAAAGA-3′.
In vitro and in vivo FRAP assays
The FRAP experiments were performed on a Zeiss LSM880 confocal microscope system. For FRAP of SARS2-NP, spots of approximately 2 μm in diameter in droplets of approximately 10 μm were photobleached with 20% laser power for 1 s using 488 nm lasers. Cellular FRAP was performed on a Zeiss LSM880 confocal microscope system at 37 °C in a live-cell-imaging chamber and the SARS2-NP was fully or partially photobleached with 20% laser power for 2 s using a 488 nm laser. Time-lapse images were acquired over 1 min after bleaching. Images were processed by ImageJ and FRAP data were fitted to a single exponential model using GraphPad Prism 8.
SPR
Biotin-labelled RNA was immobilized to a SA sensor chip using a Biacore T200 system (GE Healthcare). The running buffer contained 10 mM HEPES, pH 7.4, 150 mM NaCl, 3 mM EDTA and 0.05% P20. Serial dilutions of purified recombinant SARS2-NP were injected at a range of concentrations as indicated in the figure legends. The resulting data were fit to a 1:1 binding model using the Biacore Evaluation Software (GE Healthcare).
The biotin-labelled SARS-CoV-2 RNA had the following sequence: 5′-CACUCGCUAUGUCGAUAACAACUUCUGUGGCCCUGAUGGCUACCCUCUUGAGUGCAUUAAAGA-3′.
Oligo pull-down assay
To assay RNA binding to SARS2-NP in cell lysates, HEK293T cells were transfected with the indicated plasmids. After 36 h, the HEK293T cells were washed twice, collected and lysed in buffer containing 50 mM Tris–HCl, 150 mM NaCl, 1% NP-40, 5 mM EDTA, protease inhibitor cocktail and phosphatase inhibitor cocktail (Roche) for 30 min. The cell debris was pelleted by centrifugation at 16,000g for 15 min at 4 °C. Biotin-labelled RNA (2 μg) was first immobilized onto 15 μl of streptavidin sepharose beads. The beads were washed three times with lysis buffer (described above) to remove unconjugated ligands and then added to pre-cleared lysates to perform the standard pull-down assay.
Interfering peptides
The NIPs were as follows: NIP-I, H-FNGQTQEPGRpprrrqrrkkrg-OH (molecular weight (Mw), 2,706; lowercase font, HIV-TAT sequence); NIP-II, H-IRSMGFFASASPAFQAIQPWpprrrqrrkkrg-OH (Mw, 3,785); NIP-III, H-LKIAGTYTLWTGSPTVEMGIpprrrqrrkkrg-OH (Mw, 3,711); NIP-IV, H-SDASSMSQQLQKSFDDLpprrrqrrkkrg-OH (Mw, 3,460); and NIP-V, H-IHKNLLIVQDKFpprrrqrrkkrg-OH (Mw, 3,041).
The interfering peptides were manufactured by GL Biochem at >99% purity and stored at −20 °C in powder aliquots of 1 mg until use to avoid freeze–thaw artefacts. For in vitro experiments, the immunoprecipitates were dissolved in PBS to generate a 25 mM stock. For in vivo experiments, the immunoprecipitates were dissolved in PBS to generate a 5 mg ml−1 stock solution, which was kept on ice until injection. Before injection, the solution was brought to room temperature.
Viral infection and plaque assay
To construct the recombinant VSV carrying GFP, SARS2-NP WT or SARS2-NP d_DD, the coding sequence of the GFP, SARS2-NP WT or SARS2-NP d_DD was inserted into VSV backbones between the VSV glycoprotein (G protein) and polymerase protein (L protein) using NheI and XhoI restriction enzyme sites30. The VSV backbone harbours the M51R substitution in the matrix (M) gene, attenuating the virulence of original strain28,29. The recombinant virus (GFP–VSV, NP–VSV or NP–VSV d_DD) was then recovered in Vero E6 cells30. Mouse macrophages or other cells (2 × 105) were plated 24 h before infection. The cells were infected with VSV (m.o.i. of 0.1) or SeV (100 haemagglutination units (HAU) ml−1) for varying times as indicated in the figures. VSV plaque assay and VSV replication were determined by a standard TCID50 assay on permissive Vero E6 cell monolayers in 96-well plates with a tenfold dilution series of the samples. After 1 h of infection, the plates were incubated for 48 h. The medium was then removed, and the cells were fixed with 4% paraformaldehyde for 15 min and stained with 1% crystal violet for 30 min before plaque counting.
Viral infection in vivo
For in vivo viral infection studies, eight-week-old mice were infected with VSV (5 × 108 p.f.u. per mouse) by i.p. injection for 24 h. The eight-week-old hACE2 transgenic mice were intranasally inoculated with SARS-CoV-2 strain HB-01 at a dosage of 1 × 105 TCID50 for 16 and 24 h. We collected blood from the orbital sinus for ELISA and the lungs, spleen and liver from each mouse for analysis of RNA, protein and viral titres. To measure the VSV titres in the lungs, spleen and liver, snap-frozen tissues were weighed and homogenized three times (5 s each) in MEM. After homogenization, the suspensions were centrifuged at 1,620g for 30 min and the supernatants were used for plaque assays on monolayers of Vero E6 cells seeded in 96-well plates with a tenfold dilution series of the samples. For the survival experiments, the mice were monitored for survival after infection with VSV.
ELISA
The concentrations of IFN-β in culture supernatants and serum were measured using ELISA kits (R&D Systems).
Lung histology
Lungs from control or virus-infected mice were dissected, fixed in 10% phosphate-buffered formalin, embedded in paraffin, sectioned, stained with H&E solution and examined by light microscopy for histological changes.
Immunohistochemical staining and evaluation
Primary antibodies to Spike (Abcam, ab273433; 1:1,000) and Spike (ABclonal, A20022; 1:1,000) were used for immunohistochemical staining of formalin-fixed paraffin-embedded microarrays of lung tissues from each group of mice, according to previously described staining protocols77. The quantification of staining was expressed as an H score. The H score was determined using the formula 3 × the percentage of strongly stained cells + 2 × the percentage of moderately stained cells + the percentage of weakly stained cells, yielding a range of 0–300.
Statistics and reproducibility
Statistical analyses were performed using a two-tailed unpaired Student’s t-test or as indicated in the figure legends. All of the assays were performed three times with similar results. The exact value of n, representing the number of mice in the experiments, is indicated in the figure legends. Statistical differences with a P value of 0.05 or less were considered significant. There was no exclusion of data points or mice. No randomization or blinding were used.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. All other raw data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Huang, C. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395, 497–506 (2020).
Wu, A. et al. Genome composition and divergence of the novel coronavirus (2019-nCoV) originating in China. Cell Host Microbe 27, 325–328 (2020).
Artika, I. M., Dewantari, A. K. & Wiyatno, A. Molecular biology of coronaviruses: current knowledge. Heliyon 6, e04743 (2020).
Paces, J., Strizova, Z., Smrz, D. & Cerny, J. COVID-19 and the immune system. Physiol. Res. 69, 379–388 (2020).
McKechnie, J. L. & Blish, C. A. The innate immune system: fighting on the front lines or fanning the flames of COVID-19? Cell Host Microbe 27, 863–869 (2020).
Li, G. et al. Coronavirus infections and immune responses. J. Med. Virol. 92, 424–432 (2020).
Zalinger, Z. B., Elliott, R., Rose, K. M. & Weiss, S. R. MDA5 is critical to host defense during infection with murine coronavirus. J. Virol. 89, 12330–12340 (2015).
Li, J., Liu, Y. & Zhang, X. Murine coronavirus induces type I interferon in oligodendrocytes through recognition by RIG-I and MDA5. J. Virol. 84, 6472–6482 (2010).
Roth-Cross, J. K., Bender, S. J. & Weiss, S. R. Murine coronavirus mouse hepatitis virus is recognized by MDA5 and induces type I interferon in brain macrophages/microglia. J. Virol. 82, 9829–9838 (2008).
Totura, A. L. et al. Toll-like receptor 3 signaling via TRIF contributes to a protective innate immune response to severe acute respiratory syndrome coronavirus infection. mBio 6, e00638-15 (2015).
Kadowaki, N., Antonenko, S., Lau, J. Y. & Liu, Y. J. Natural interferon α/β-producing cells link innate and adaptive immunity. J. Exp. Med. 192, 219–226 (2000).
Brubaker, S. W., Bonham, K. S., Zanoni, I. & Kagan, J. C. Innate immune pattern recognition: a cell biological perspective. Annu. Rev. Immunol. 33, 257–290 (2015).
Fischer, S. Pattern recognition receptors and control of innate immunity: role of nucleic acids. Curr. Pharm. Biotechnol. 19, 1203–1209 (2018).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Hurst, K. R., Ye, R., Goebel, S. J., Jayaraman, P. & Masters, P. S. An interaction between the nucleocapsid protein and a component of the replicase-transcriptase complex is crucial for the infectivity of coronavirus genomic RNA. J. Virol. 84, 10276–10288 (2010).
Ye, Q., West, A. M. V., Silletti, S. & Corbett, K. D. Architecture and self-assembly of the SARS-CoV-2 nucleocapsid protein. Protein Sci. 29, 1890–1901 (2020).
Peng, Y. et al. Structures of the SARS-CoV-2 nucleocapsid and their perspectives for drug design. EMBO J. 39, e105938 (2020).
Zhou, R., Zeng, R., von Brunn, A. & Lei, J. Structural characterization of the C-terminal domain of SARS-CoV-2 nucleocapsid protein. Mol. Biomed. 1, 2 (2020).
Hyman, A. A., Weber, C. A. & Julicher, F. Liquid–liquid phase separation in biology. Annu. Rev. Cell Dev. Biol. 30, 39–58 (2014).
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters. 3, e201702000010 (2017).
Soderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Soderberg, O. et al. Characterizing proteins and their interactions in cells and tissues using the in situ proximity ligation assay. Methods 45, 227–232 (2008).
Chang, C. K. et al. Multiple nucleic acid binding sites and intrinsic disorder of severe acute respiratory syndrome coronavirus nucleocapsid protein: implications for ribonucleocapsid protein packaging. J. Virol. 83, 2255–2264 (2009).
Carlson, C. R. et al. Phosphoregulation of phase separation by the SARS-CoV-2 N protein suggests a biophysical basis for its dual functions. Mol. Cell 80, 1092–1103 (2020).
Jayaram, H. et al. X-ray structures of the N- and C-terminal domains of a coronavirus nucleocapsid protein: implications for nucleocapsid formation. J. Virol. 80, 6612–6620 (2006).
Ahmed, M. & Lyles, D. S. Identification of a consensus mutation in M protein of vesicular stomatitis virus from persistently infected cells that affects inhibition of host-directed gene expression. Virology 237, 378–388 (1997).
Desforges, M. et al. Different host-cell shutoff strategies related to the matrix protein lead to persistence of vesicular stomatitis virus mutants on fibroblast cells. Virus Res. 76, 87–102 (2001).
Lawson, N. D., Stillman, E. A., Whitt, M. A. & Rose, J. K. Recombinant vesicular stomatitis viruses from DNA. Proc. Natl Acad. Sci. USA 92, 4477–4481 (1995).
Dragan, A. I., Hargreaves, V. V., Makeyeva, E. N. & Privalov, P. L. Mechanisms of activation of interferon regulator factor 3: the role of C-terminal domain phosphorylation in IRF-3 dimerization and DNA binding. Nucleic Acids Res. 35, 3525–3534 (2007).
Panne, D., McWhirter, S. M., Maniatis, T. & Harrison, S. C. Interferon regulatory factor 3 is regulated by a dual phosphorylation-dependent switch. J. Biol. Chem. 282, 22816–22822 (2007).
Hou, F. et al. MAVS forms functional prion-like aggregates to activate and propagate antiviral innate immune response. Cell 146, 448–461 (2011).
Liu, B. et al. The ubiquitin E3 ligase TRIM31 promotes aggregation and activation of the signaling adaptor MAVS through Lys63-linked polyubiquitination. Nat. Immunol. 18, 214–224 (2017).
King, O. D., Gitler, A. D. & Shorter, J. The tip of the iceberg: RNA-binding proteins with prion-like domains in neurodegenerative disease. Brain Res. 1462, 61–80 (2012).
Dai, T. et al. FAF1 regulates antiviral immunity by inhibiting MAVS but is antagonized by phosphorylation upon viral infection. Cell Host Microbe 24, 776–790 (2018).
Ryu, Y. & Schultz, P. G. Efficient incorporation of unnatural amino acids into proteins in Escherichia coli. Nat. Methods 3, 263–265 (2006).
Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Genetically encoding Nε-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Savastano, A., Ibanez de Opakua, A., Rankovic, M. & Zweckstetter, M. Nucleocapsid protein of SARS-CoV-2 phase separates into RNA-rich polymerase-containing condensates. Nat. Commun. 11, 6041 (2020).
Zhao, W. M. et al. The 2019 novel coronavirus resource (in Chinese). Yi Chuan 42, 212–221 (2020).
Gong, Z. et al. An online coronavirus analysis platform from the national genomics data center. Zool. Res. 41, 705–708 (2020).
Guichard, G. et al. Antigenic mimicry of natural L-peptides with retro-inverso-peptidomimetics. Proc. Natl Acad. Sci. USA 91, 9765–9769 (1994).
Baar, M. P. et al. Targeted apoptosis of senescent cells restores tissue homeostasis in response to chemotoxicity and aging. Cell 169, 132–147 (2017).
Yuan, Y. et al. Targeting UBE4A revives viperin protein in epithelium to enhance host antiviral defense. Mol. Cell 77, 734–747 (2020).
Borsello, T. et al. A peptide inhibitor of c-Jun N-terminal kinase protects against excitotoxicity and cerebral ischemia. Nat. Med. 9, 1180–1186 (2003).
Beydoun, T. et al. Subconjunctival injection of XG-102, a JNK inhibitor peptide, in patients with intraocular inflammation: a safety and tolerability study. J. Ocul. Pharm. Ther. 31, 93–99 (2015).
Suckfuell, M. et al. Efficacy and safety of AM-111 in the treatment of acute sensorineural hearing loss: a double-blind, randomized, placebo-controlled phase II study. Otol. Neurotol. 35, 1317–1326 (2014).
Deloche, C. et al. XG-102 administered to healthy male volunteers as a single intravenous infusion: a randomized, double-blind, placebo-controlled, dose-escalating study. Pharm. Res Perspect. 2, e00020 (2014).
Warso, M. A. et al. A first-in-class, first-in-human, phase I trial of p28, a non-HDM2-mediated peptide inhibitor of p53 ubiquitination in patients with advanced solid tumours. Br. J. Cancer 108, 1061–1070 (2013).
Herce, H. D. & Garcia, A. E. Molecular dynamics simulations suggest a mechanism for translocation of the HIV-1 TAT peptide across lipid membranes. Proc. Natl Acad. Sci. USA 104, 20805–20810 (2007).
Falzarano, D. et al. Inhibition of novel β coronavirus replication by a combination of interferon-α2b and ribavirin. Sci. Rep. 3, 1686 (2013).
Chan, J. F. et al. Treatment with lopinavir/ritonavir or interferon-β1b improves outcome of MERS-CoV infection in a nonhuman primate model of common marmoset. J. Infect. Dis. 212, 1904–1913 (2015).
Sheahan, T. P. et al. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat. Commun. 11, 222 (2020).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 (2020).
Chan, R. W. et al. Tropism of and innate immune responses to the novel human betacoronavirus lineage C virus in human ex vivo respiratory organ cultures. J. Virol. 87, 6604–6614 (2013).
Cinatl, J. et al. Treatment of SARS with human interferons. Lancet 362, 293–294 (2003).
Stroher, U. et al. Severe acute respiratory syndrome-related coronavirus is inhibited by interferon-α. J. Infect. Dis. 189, 1164–1167 (2004).
de Wilde, A. H. et al. MERS-coronavirus replication induces severe in vitro cytopathology and is strongly inhibited by cyclosporin A or interferon-α treatment. J. Gen. Virol. 94, 1749–1760 (2013).
Lokugamage, K. G. et al. Type I interferon susceptibility distinguishes SARS-CoV-2 from SARS-CoV. J. Virol. 94, e01410-20 (2020).
Chu, H. et al. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin. Infect. Dis. 71, 1400–1409 (2020).
Chan, J. F. et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 9, 221–236 (2020).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
de Wit, E., van Doremalen, N., Falzarano, D. & Munster, V. J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 14, 523–534 (2016).
Mesel-Lemoine, M. et al. A human coronavirus responsible for the common cold massively kills dendritic cells but not monocytes. J. Virol. 86, 7577–7587 (2012).
Donnelly, C. A. et al. Epidemiological determinants of spread of causal agent of severe acute respiratory syndrome in Hong Kong. Lancet 361, 1761–1766 (2003).
Chan, J. W. M. et al. Short term outcome and risk factors for adverse clinical outcomes in adults with severe acute respiratory syndrome (SARS). Thorax 58, 686–689 (2003).
Mizumoto, K. et al. Real-time characterization of risks of death associated with the Middle East respiratory syndrome (MERS) in the Republic of Korea, 2015. BMC Med. 13, 228 (2015).
Perdikari, T. M. et al. SARS-CoV-2 nucleocapsid protein phase-separates with RNA and with human hnRNPs. EMBO J. 39, e106478 (2020).
Chen, H. et al. Liquid–liquid phase separation by SARS-CoV-2 nucleocapsid protein and RNA. Cell Res. 30, 1143–1145 (2020).
Iserman, C. et al. Genomic RNA elements drive phase separation of the SARS-CoV-2 nucleocapsid. Mol. Cell 80, 1078–1091 (2020).
Zhao, M. et al. GCG inhibits SARS-CoV-2 replication by disrupting the liquid phase condensation of its nucleocapsid protein. Nat. Commun. 12, 2114 (2021).
Wu, Y. et al. RNA-induced liquid phase separation of SARS-CoV-2 nucleocapsid protein facilitates NF-κB hyper-activation and inflammation. Signal Transduct. Target Ther. 6, 167 (2021).
Cubuk, J. et al. The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA. Nat. Commun. 12, 1936 (2021).
Wang, S. et al. YAP antagonizes innate antiviral immunity and is targeted for lysosomal degradation through IKKɛ-mediated phosphorylation. Nat. Immunol. 18, 733–743 (2017).
Zhang, L. et al. TRAF4 promotes TGF-β receptor signaling and drives breast cancer metastasis. Mol. Cell 51, 559–572 (2013).
Zhou, F. et al. Nuclear receptor NR4A1 promotes breast cancer invasion and metastasis by activating TGF-β signalling. Nat. Commun. 5, 3388 (2014).
Acknowledgements
This work was supported by a special programme from the Ministry of Science and Technology of China (grant no. 2016YFA0502500 to L.Z.), the Chinese National Natural Science Funds (grant nos U20A201376, 82041009, 31871405, 91753139, 31925013, 31671457, 31571460, 31870902, 32070907, 31871405, 31771619, 92053114, and 32070632), National Postdoctoral Program for Innovative Talent (grant no. BX2021208), Jiangsu National Science Foundation (grant nos BK20180043 and 19KJA550003), Shenzhen Science and Technology Program (grant no. JSGG20200225150431472), and special COVID-19 program of the Sino-German Center for Research Promotion (grant no. C-0023).
Author information
Authors and Affiliations
Contributions
S.W., T.D. and Z.Q. designed the experiments and analysed the data. S.W., T.D., Z.Q., F.C., L.L. and T.P. performed the experiments. B.Y. performed the mass spectrometry analysis. H.H., H.L. and L.Z. provided valuable discussion. T.D. and F.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The peptide NIP-V are subject to patent applications (patent no. 202110466103.7; China; F.Z., T.D., S.W., and L.Z.) and may be shared with research organizations for research and educational purpose only under an material transfer agreement to be discussed in good faith with the recipient; such a material transfer agreement may restrict the recipient to make any modifications to these materials. All of the other authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Luke O’Neill and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SARS2-NP undergoes LLPS.
a, Domain structure (upper) and the intrinsically disordered tendency (lower) of SARS2-NP. IUPred2, ANCHOR2, PrD.like and VL-XT assigned scores of disordered tendencies between 0 and 1 to the sequences were shown. b, Bacterially purified SARS2-NP wt, d_IDR1, d_IDR2, d_IDR3, d_RBD, and d_DD proteins were analysed by SDS–PAGE and detected by Coomassie blue staining. c-e, 5 μM GFP–SARS2-NP were treated with 5% Hex (c), heated-inactivated (5 min at 95 °C and immediately put on ice for 5 min) (d), or treated with 100 μg/ml Proteinase K for 30 min at 40 °C (e), and then subjected to droplet formation assay in vitro (150 mM NaCl, pH 5.5, room temperature). Scale bar, 10 μm. Mean ± s.d., n = 6 independent experiments. f, SDD–AGE analysis of SARS2-NP aggregation (top) and SDS–PAGE (bottom) of cytosolic lysate of HEK293T cells transfected with plasmids expressing Myc–SARS2-NP wt and treated with increased β-mercaptoethanol as indicated. g, SDD–AGE analysis of SARS2-NP aggregation (top) and SDS–PAGE (bottom) of cytosolic lysate of HEK293T cells transfected with plasmids expressing Myc–SARS2-NP wt or d_DD. h, Sucrose-gradient ultracentrifugation analysis of total cell lysates of HEK293T cells transfected with Myc–SARS2-NP wt or d_DD plasmids. The data are representative of three independent experiments (b, f-h). Statistical analysis was performed using two-tailed Student’s t-test (c-e). Source data are available online.
Extended Data Fig. 2 SARS2-NP dampens IFN-β signalling relies on its DD.
a, IFN-β-Luc (left) and PRDs I–III-Luc (right) activity in HEK293T cells transfected with Co.vec, SARS2-NP wt or truncated plasmids, followed by infection with SeV for 12 h. b, qPCR analysis of CCL5 (left) and CXCL10 (right) mRNA in HEK293T cells transfected with Co.vec, SARS2-NP wt or truncated plasmids, followed by infection with SeV for 12 h. c, qPCR analysis of IFNB1 mRNA in A549 transfected with Co.vec, SARS2-NP wt or truncated plasmids, followed by infection with SeV for 6 h. d, qPCR analysis of IFNB1 mRNA in A549 transfected with Co.vec, SARS2-NP wt or d_DD plasmids, followed by stimulation with Poly (I:C) or 5’-ppp RNA for 6 h. e, qPCR analysis of IFNB1 mRNA (left), VSV copy number (middle), and plaque assay of VSV (right) in A549 transfected with Co.vec, SARS2-NP wt or d_DD plasmids, followed by infected with VSV (m.o.i., 0.1) for 12 h. f, IB analysis of VSV-G in A549 transfected with Co.vec, SARS2-NP wt or d_DD plasmids, followed by infected with VSV (m.o.i., 0.1) for 12 h. Quantification of the VSV-G band intensities were shown below. g, Left: SeV-induced CPE in MEF cells transfected with Co.vec, SARS2-NP wt or d_DD plasmids, followed by infected for various times with SeV. Right: CPE rates shown was quantified by ImageJ. Mean ± s.d., n = 3 biological independent samples (a-e, f (lower), g (right)). Statistical analysis was performed using two-tailed Student’s t-test (a-e, f (lower)) or two-way ANOVA (g (right)). Source data are available online.
Extended Data Fig. 3 SARS2-NP wt, but not d_DD, negatively regulates MAVS.
a, a schematic diagram of MAVS truncations. b, Immunofluorescence microscopy and DAPI staining of U2OS cells transfected with MAVS together with Co.vec, SARS2-NP wt or d_DD plasmids, followed by infection for 8 h with SeV. Scale bar, 10 μm. Confocal images represent five cells with a similar pattern. c, Upper: GFP–MAVS (green) and mCherry–NP d_DD (magenta) (200 nM each) were mixed with the indicated module concentration and were imaged for fluorescence. The data are representative of three independent experiments. Scale bar, 10 μm. Lower: PCs (means of duplicate samples) of GFP–MAVS and mCherry–NP wt or d_DD into droplets formed by the indicated module concentrations of MAVS and NP deletions. Source data are available online.
Extended Data Fig. 4 The acetylation of SARS2-NP on Lys375 reduces LLPS and its ability to bind RNA.
a, IB of TCLs and immunoprecipitants derived from HEK293T cells transfected with Myc–SARS2-NP and HA-CBP, p300, PCAF, GCN5, KAT5, or KAT8 plasmids. b, IB of TCLs and proteins immunoprecipitated, with antibody specifically recognizing acetylated-Lys375 of SARS2-NP, from HEK293T cells transfected with expression plasmids for Myc–SARS2-NP wt or K375R mutant along with or without HA-CBP. c, IB of TCLs and proteins immunoprecipitated, with antibody specifically recognizing acetylated-Lys375 of SARS2-NP, from HEK293T cells transfected with Myc–SARS2-NP and control small interfering RNA (siRNA) or three independent CBP siRNA (#1, #2 and #3). d, IB of TCL and immunoprecipitants derived from HEK293T cells transfected with Myc–SARS2-NP and HA-CBP, and then treated with or without DMSO, TSA (5 μM), NAM (10 mM) as indicated for 6 h. e, IB of TCLs and immunoprecipitants derived from HEK293T cells transfected with Myc–SARS2-NP and HA-CBP, along with Flag-HADC1-11 plasmids (left) or Sirt1–7 plasmids (right). f, IB analysis of TCL and streptavidin RNA pull-down derived from HEK293T cells transfected with Myc–SARS2-NP along with or without CBP plasmid. Quantification of the binding Myc–SARS2-NP band intensities were shown right (mean ± s.d.). g, IB analysis of TCL and streptavidin RNA pull-down derived from HEK293T cells transfected with Myc–SARS2-NP along with or without CBP siRNA. Quantification of the binding Myc–SARS2-NP band intensities were shown right (mean ± s.d.). h, IFN-β-Luc activity in HEK293T cells transfected with the reporter and SARS2-NP wt, K375Q, or K375R plasmids, followed by stimulated with or without SeV (left), VSV (middle), or MAVS (right). i, VSV–GFP intensity of HEK293T cells transfected with Co.vec, SARS2-NP wt, K375Q, or K375R plasmids, followed by infection for 12 h with GFP–VSV, was quantified by ImageJ. Blots are representative of three independent experiments (a-g). Data are presented as mean ± s.d., n = 3 biological independent samples (h, i). Statistical analysis was performed using two-tailed Student’s t-test (f (right), g (right), h, i). Source data are available online.
Extended Data Fig. 5 Efficiency of SARS2-NP wt and K375 mutants in disrupting MAVS droplets formation.
Left: GFP–MAVS (green), mCherry–NP wt or mutants (magenta), and RNA (200 nM each) were mixed with the indicated module concentration and were imaged for fluorescence. The data are representative of three independent experiments. Scale bar, 5 μm. Right: PCs (means of duplicate samples) of GFP–MAVS and mCherry–NP wt or mutants into droplets formed by the indicated module concentrations of MAVS and NP mutants. Source data are available online.
Extended Data Fig. 6 K375N and K375E variants that mimic SARS2-NP acetylation exhibit lower levels of LLPS and reduced ability to antagonize innate antiviral immunity.
a, SDD–AGE analysis of SARS2-NP aggregation (top) and SDS–PAGE (bottom) of cytosolic lysate of HEK293T cells transfected with plasmids expressing Myc–SARS2-NP wt, K375E, or K375N plasmids. b, IFN-β-Luc activity in HEK293T cells transfected with the reporter and SARS2-NP wt, K375N, or K375E plasmids, followed by stimulated with or without SeV (left), VSV (middle), or MAVS (right). c, qPCR analysis of IFNB1 mRNA in HEK293T cells transfected with Co.vec, SARS2-NP wt, K375N, or K375E plasmids, followed by infection with SeV or VSV for 12 h. d, VSV–GFP intensity of HEK293T cells transfected with Co.vec, SARS2-NP wt, K375N, or K375E plasmids, followed by infection for 12 h with GFP–VSV, was quantified by ImageJ. Blots are representative of three independent experiments (a). Data are presented as mean ± s.d., n = 3 biological independent samples. Statistical analysis was performed using two-tailed Student’s t-test (b–d). Source data are available online.
Extended Data Fig. 7 NIP-V peptide targeting the DD efficiently disrupts SARS2-NP LLPS and thus elevates the innate antiviral responses in vitro.
a, Related to Fig. 8c: statistical analysis of the droplet formation of GFP–SARS2-NP (10 μM) mixed with control PBS or NIPs (5 µM) (mean ± s.d., n = 6 independent experiments). b, Related to Fig. 8d: the HeLa cells transfected with GFP–SARS2-NP and then treated with NIPs (50 µM) for 12 h. The cells harbouring NP droplets were quantified (mean ± s.d., n = 3 independent experiments). c, IB of TCL and immunoprecipitants derived from HEK293T cells transfected with Flag- and Myc–SARS2-NP, and treated with or without 200 µM NIP-V for 12 h. d, qPCR analysis of IFNB1 mRNA in HEK293T cells transfected with RIG-IN, MAVS, or IRF3-5D along with SARS2-NP, and treated with 50 µM or 100 µM NIP-V for 12 h as indicated (mean ± s.d., n = 3 biological independent samples). Statistical analysis was performed using two-tailed Student’s t-test (a, b, d). Blots are representative of three independent experiments (c). Source data are available online.
Extended Data Fig. 8 NIP-V peptide targeting DD elevates the innate antiviral responses against NP–VSV and SARS-CoV-2 viruses in vivo.
a-b, qPCR analysis of Ifnb1, Isg56 (a) and Cxcl10 (b) mRNA in the spleen, liver and lung of mice (n = 5 mice per group) injected with PBS (i.p.) or NIP-V (5 mg/kg, i.p.) for 2 h and then infected with NP–VSV (5 × 108 PFU per mouse, i.p.) for 24 h. c, ELISA determination of IFN-β level in serum from mice as in a. d-e, qPCR analysis of VSV RNA (d) and plaque assay of VSV (e) in the spleen, liver, and lung of infected mice as in a. f, IB analysis of VSV-G in the spleen (top), liver (middle) and lung (bottom) of infected mice selected randomly as in a. n = 3 mice per group. g, Survival curves of mice (n = 10 mice per group) administrated with PBS or NIP-V (5 mg/kg, i.p.) for 2 h and then infected with NP–VSV (1 × 109 PFU per mouse, i.p.). h, Microscopy of hematoxylin-and-eosin-stained lung sections from mice treated with PBS or VSV as in a. Scale bar, 500 µm (top) or 100 µm (bottom). Images represent five independent samples with a similar pattern. i, qPCR of Isg56 mRNA in the spleen, liver and lung (right) of hACE2 transgenic mice (n = 6 mice per control uninfected groups, n = 8 mice per infected groups) pretreated with PBS or NIP-V (25 mg/kg, i.p.) for 2 h and then given intranasal inoculation with SARS-CoV-2 (1×105 TCID50 per mouse) for another 16 or 24 h. j, Immunofluorescence assay of the SARS2-NP antigen (green) of lung sections from hACE2 transgenic mice pretreated with PBS or NIP-V (25 mg/kg, i.p.) for 2 h and then intranasally inoculated with SARS-CoV-2 (1×105 TCID50 per mouse) for 24 h. The nuclei were stained with DAPI (blue). Scale bar, 500 µm. n = 8 independent samples. Data are presented as mean ± s.d. (a-e, i, j (right)). Statistical analysis was performed using two-tailed Student’s t-test (a-e, i, j (right)) or log-rank test (g). Source data are available online.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unmodified blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unmodified blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unmodified blots.
Source Data Fig. 4
Unmodified blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unmodified blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unmodified blots.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unmodified blots.
Source Data Fig. 8
Statistical source data.
Source Data Fig. 8
Unmodified blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unmodified blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unmodified blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unmodified blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unmodified blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unmodified blots.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unmodified blots.
Rights and permissions
About this article
Cite this article
Wang, S., Dai, T., Qin, Z. et al. Targeting liquid–liquid phase separation of SARS-CoV-2 nucleocapsid protein promotes innate antiviral immunity by elevating MAVS activity. Nat Cell Biol 23, 718–732 (2021). https://doi.org/10.1038/s41556-021-00710-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00710-0
This article is cited by
-
Immunological mechanisms of the nucleocapsid protein in COVID-19
Scientific Reports (2024)
-
The E3 ubiquitin ligase MARCH2 protects against myocardial ischemia-reperfusion injury through inhibiting pyroptosis via negative regulation of PGAM5/MAVS/NLRP3 axis
Cell Discovery (2024)
-
SARS-CoV-2 biology and host interactions
Nature Reviews Microbiology (2024)
-
TRIM28-mediated nucleocapsid protein SUMOylation enhances SARS-CoV-2 virulence
Nature Communications (2024)
-
The SARS-CoV-2 nucleocapsid protein: its role in the viral life cycle, structure and functions, and use as a potential target in the development of vaccines and diagnostics
Virology Journal (2023)