Abstract
BALB/c mice heterozygous for Trp53 develop a high proportion of spontaneous mammary tumors, a phenotype distinct from other mouse strains. BALB/c-Trp53+/− female mice, thus, resemble the hereditary Li-Fraumeni syndrome (LFS) characterized by early-onset of breast cancer, even though LFS involves TP53 mutations, which may involve not only loss- but also gain-of-function. Previous analysis of tumors in BALB/c-Trp53+/− females showed frequent loss of heterozygosity involving the wild-type allele of Trp53 and displayed characteristics indicative of mitotic recombination. Critical involvement of DNA double-strand break (DSB) repair dysfunction, particularly of homologous recombination (HR), was also noticed in the etiology of human breast cancer. To better define functional alterations in BALB/c-Trp53+/− mice, we applied a fluorescence-based DSB repair assay on mouse embryonic fibroblasts (MEFs) from BALB/c-Trp53+/− versus C57BL/6J-Trp53+/− mice. This approach revealed deregulation of HR but not non-homologous end-joining (NHEJ) in BALB/c-Trp53+/−, which was further confirmed for mammary epithelial cells. Screening of a small interfering RNA-library targeting DSB repair, recombination, replication and signaling genes, identified 25 genes causing differences between homologous DSB repair in the two strains upon silencing. Interactome analysis of the hits revealed clustering of replication-related and fanconi anemia (FA)/breast cancer susceptibility (BRCA) genes. Further dissection of the functional change in BALB/c-Trp53+/− by immunofluorescence microscopy of nuclear 53BP1, Replication protein A (RPA) and Rad51 foci uncovered differences in crosslink and replication-associated repair. Chromosome breakage, G2 arrest and biochemical analyses indicated a FA pathway defect downstream of FancD2 associated with reduced levels of BRCA2. Consistent with polygenic models for BRCA, mammary carcinogenesis in BALB/c-Trp53+/− mice may, therefore, be promoted by a BRCA modifier allele in the FA pathway in the context of partial p53 loss-of-function.
Similar content being viewed by others
Introduction
Cells from Li-Fraumeni syndrome (LFS) patients have been shown to accumulate chromosome instabilities.1 More recently, high-resolution genome-wide Single Nucleotide Polymorphism (SNP)-chip-analysis revealed excessive copy number variations, particularly loss of heterozygosity (LOH), at various loci in the genome of peripheral blood lymphocytes among carriers of germline TP53 mutations with a further increase in mutation carriers affected with cancer.2 Copy number variations occur 100–10 000 times more frequently than point mutations in the human genome, and are, therefore, particularly relevant for tumorigenesis.3 Non-allelic HR processes give rise to copy number variations, which is consistent with observations in mice and mouse embryonic fibroblasts (MEFs), indicating that p53 is haploinsufficient for suppression of mitotic recombination events.4, 5 Biochemical and cell-based studies further demonstrated that p53 suppresses HR, particularly between short stretches of homologies, thereby causing a shift to low-fidelity processes upon inactivation.6, 7 Inherited mutations in DNA double-strand break (DSB) repair genes that predispose to breast cancer (for example, BRCA1, BRCA2/FANCD1, BRIP1/FANCJ, PALB2/FANCN and RAD51C/FANCO) identify the vulnerability of this pathway in breast carcinogenesis. Therefore, impaired suppression of HR in LFS patients appears causally linked to breast carcinogenesis, the most common tumor in women with inherited mutations in TP53.8
In mice, heterozygous mutations in the gene encoding p53 (designated Trp53) also predispose to tumors, but the prevalence of tumors differs greatly among strains. While lymphomas are prevalent regardless of genetic background, spontaneous mammary tumors are only observed in BALB/c-Trp53+/− mice.9 Polymorphisms in Prkdc that participates in NHEJ and the cell cycle regulator Cdkn2a interact and contribute to differences in the spectrum of tumors.10 Among the strain differences is the greatly increased incidence of LOH at Trp53 in tumors from BALB/c-Trp53+/− mice but rarity in C57BL/6-Trp53+/− mice.11 The genetic predisposition to mammary tumors has been mapped to two distinct loci on chromosome 7, as well as a recessive-acting locus on the X-chromosome.12, 13 These results demonstrate the multigenic nature of the predisposition to mammary tumors that interact with p53 deficiency. Conversely, the pathways present in C57BL/6 mice compensate for the haploinsufficiency in Trp53 rendering mice resistant to mammary tumors, but not tumors in other tissues.
Given the prominent role of DSB repair in heritable breast cancer, these pathways provide a plausible target for the differences in susceptibility to mammary tumors observed among strains of rodents.14 As mutations in the p53 pathway remain one of the most common genetic alteration in sporadic breast cancers in women,15 pathways that interact with p53 and can complement p53 insufficiency would be valuable therapeutic targets. Therefore, a small interfering RNA (siRNA) screen of DSB repair pathways was undertaken to examine potential differences in DNA repair among C57BL/6 and BALB/c mice in the context of haploinsufficiency for p53. The results identify key targets within fanconi anemia (FA) and BRCA complexes, as well as genes involved in DNA replication and repair. Although these functional clusters contain many genes, the siRNA screen identifies those that are most likely rate-limiting, and thus, most sensitive to disruption or to therapies to restore function. Functional assays demonstrated delays in the processing of DSBs in BALB/c mice that are similar to those observed in FA patient cells. Therefore, these results identify interactions between p53 loss and low-penetrance defects in the FA pathway, which predispose to breast cancer.
Results
Comparative DSB repair analysis
To understand whether early-onset of mammary tumors in BALB/c-Trp53+/− mice is associated with DSB repair deregulation as had been observed in primary cells from individuals with hereditary breast cancer risk,16 we compared DSB repair in primary cells derived from BALB/c-Trp53+/− and C57BL/6-Trp53+/− mice. To this end we transfected MEFs with EGFP-based DSB repair substrates comprising a recognition site for endonuclease I-SceI, together with I-SceI expression plasmid for targeted cleavage. We applied substrates EJ-EGFP and ΔEGFP/3′EGFP enabling detection of repair changes in high-risk individuals.16 In substrate EJ-EGFP, the I-SceI site is flanked by 5 bp microhomologies, enabling in-frame reconstitution of EGFP by microhomology-mediated NHEJ (Figures 1a and b). Δ-EGFP/3′EGFP was designed for analysis of homologous repair between internally mutated Δ-EGFP and 5′-mutated 3′EGFP. When monitoring EGFP-reconstitution by fluorescence activated cell sorting (FACS) analysis (normalized with transfection efficiencies, see Materials and methods), we observed that homologous repair was twofold (P=0.0049–0.0057) more active in BALB/c-Trp53+/− than C57BL/6-Trp53+/− MEFs, whereas mean NHEJ frequencies showed no significant difference (P=0.8421, see Figure 1a, Supplementary Figure 1A). As homologous DSB repair frequencies did not show statistically significant differences between C57BL/6-Trp53+/− and Trp53+/+ MEFs, but were elevated in BALB/c-Trp53+/− versus Trp53+/+ MEFs (data not shown), these results indicate that homologous DSB repair is restrained in C57BL/6 even when Trp53+/−, but derepressed in BALB/c-Trp53+/−.
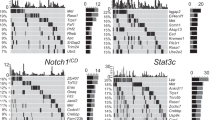
siRNA screening identifies proteins involved in differential regulation of homologous DSB repair in C57BL/6-Trp53+/− and BALB/c-Trp53+/− cells. (a) Comparison of DSB repair pathway usage in C57BL/6-Trp53+/− and BALB/c-Trp53+/− cells. C57BL/6-Trp53+/− and BALB/c-Trp53+/− MEFs were transfected with DSB repair substrate for either NHEJ (EJ-EGFP) or homologous DSB repair (Δ-EGFP/3′EGFP) and pCMV-I-SceI for targeted cleavage. EGFP-positive cells were quantified 24 h after transfection and normalized for transfection efficiencies. Mean DSB repair frequencies and SEM were calculated from 9–10 measurements each. NHEJ frequencies in C57BL/6-Trp53+/− MEFs were defined as 100% (absolute mean values, NHEJ:1 × 10−2; homologous DSB repair: 2 × 10−2). Asterisks indicate a statistically significant difference (Mann–Whitney U-test, P=0.006; **P<0.01). (b) Schematic for screening assay. Low passage BALB/c-Trp53+/− or C57BL/6-Trp53+/− MEFs were seeded on six-well plates. Twenty-four hours later they were co-transfected with a pool of four different siRNAs and a mixture of pCMV-I-SceI, repair substrate Δ-EGFP/3′EGFP comprising two inactive copies of EGFP, and filler plasmid pBS. 5′positioned Δ-EGFP contains an I-SceI restriction site (double-headed symbol) disrupting the catalytic center of EGFP. 3′positioned 3′EGFP with functional center (diamond) carries a deleterious mutation at the 5′end of the cDNA (cross). I-SceI-cleaved DSB repair substrate triggers repair via HR or SSA. Either homologous DSB repair pathway restores an active EGFP (star). The proportion of EGFP-expressing among non-fluorescent cells was measured by flow cytometry 48 h after transfection. To assess transfection efficiencies, wtEGFP expressing plasmid was added to the DNA mixture in place of filler plasmid pBS in split samples. Further split samples were subjected to LOH analysis to verify the Trp53+/− genotype of the cultures subjected to the screening rounds. (c) Validated hits of DNA repair genes in siRNA screen. Results from the siRNA screen are shown for 25 validated hits. Bars show the deviation of the target gene-specific repair frequency after knockdown relative to the repair frequency of non-silencing siRNA control siRNA-treated samples of each strain. The corresponding log2 ratios [log2(normalized DSB repair frequency BALB/c-Trp53+/−)—log2(normalized DSB repair frequency C57BL/6-Trp53+/−)] are displayed above (triangles).
BALB/c mice are moderately sensitive to ionizing radiation and show reduced repair of radiation-induced DSBs, which seems to be causally linked to two polymorphisms in the Prkdc gene encoding DNA-protein kinase (PK)cs,17 a central component of the canonical NHEJ pathway. Therefore, we investigated potential indirect effects of DNA-PK deficiency on homologous repair. For that purpose we treated BALB/c-Trp53+/− and C57BL/6-Trp53+/− MEFs with the DNA-PK inhibitor NU7441 under conditions compromising removal of radiation-induced DSBs in MEFs.18 DSB repair measurements showed that NU7441-treatment neither altered frequencies in BALB/c-Trp53+/− (P=0.1797) nor C57BL/6-Trp53+/− (P=0.9372) MEFs (Supplementary Figure 1B). For comparison, treatment with the ataxia telangiectasia mutated (ATM) and ATM and Rad3-related (ATR) inhibitor caffeine differentially affected homologous repair in MEFs from the two strains, causing a pronounced inhibition in C57BL/6-Trp53+/− (P=0.0022) compared with BALB/c-Trp53+/− cells (P=0.1320), whereas specific ATM inhibition by KU-55933 had a small effect on both C57BL/6-Trp53+/− (P=0.0022) and BALB/c-Trp53+/− cells (P=0.0411). When comparing frequencies between the strains, only caffeine treatment induced a statistically significant difference (P=0.0043). These data suggested that increased homologous repair in BALB/c-Trp53+/− MEFs is unlikely to be a consequence of the DNA-PK impairment and rather affects a pathway controlled by ATM and/or ATR.
siRNA screening and bioinformatic analysis of target genes
To identify the molecular basis for enhanced homologous repair in BALB/c-Trp53+/−, we designed a siRNA-library targeting 148 genes encompassing subgroups related to the following activities: HR, single-strand annealing (SSA), NHEJ, excision and mismatch repair, DNA damage signaling, telomere length maintenance, polymerases, helicases and nucleases (Supplementary Table 1). MEFs from BALB/c-Trp53+/− and C57BL/6-Trp53+/− mice were co-transfected with gene-specific siRNA pools (four siRNA duplexes each), substrate Δ-EGFP/3′EGFP for homologous repair (HR and SSA) and I-SceI expression plasmid (Figure 1b). LOH of the wtTrp53 allele was excluded by genomic PCR on the MEF pools. Knockdown and DSB repair in MEFs was allowed to progress for 48 h. Knockdown was verified by immunoblot analysis for two representative genes (Supplementary Figure 2A) and by Quantitative real-time PCR (qRT–PCR) for the screening hits (see Materials and methods and Supplementary Figure 2B). Additionally, siRNA pools were compared with single siRNA duplexes for five representative genes to confirm that the effect of pooled siRNAs reflected the combined effect of the corresponding four single siRNAs (Supplementary Figure 3).
Fourty-eight hours after transfection, EGFP-positive cells were quantified for calculation of DSB repair frequencies (individually normalized to transfection efficiencies determined for each siRNA pool separately). To account for day-to-day variations, we normalized screening data to our internal standard, that is, the mean frequency obtained with negative control siRNA. The results from two primary screening rounds in quadruplicates each were used to calculate differences between the mean values obtained with MEFs from BALB/c-Trp53+/− versus C57BL/6-Trp53+/− mice. Gene-specific differences reaching statistical significance (P-values <0.05 for 39 genes, see Supplementary Table 1) were validated in a rescreen, enabling selection of 25 hits with highly significant differences (P-values <0.001, Figure 1c and Table 1).
Knockdown-induced differences between the strains amounted up to ∼50% of the control values (Figure 1c). Log2 ratios of these DSB repair frequency differences were calculated as follows: log2(normalized DSB repair frequency (BALB/c-Trp53+/-))—log2(normalized DSB repair frequency (C57BL/6-Trp53+/-)), and the resulting values ranging between 0.39 and 0.75 used to assess the relative effect of the respective knockdown. Thus, in Figure 1c, the identified genes were sorted according to the differences in DSB repair frequencies (log2 ratios) caused by knockdown in BALB/c-Trp53+/− versus C57BL/6-Trp53+/− MEFs. Interestingly, even though inactivation of homologous DSB repair components is expected to impair this pathway, downregulation of critical components such as Xrcc2 or Palb2 did not cause a decrease in homologous DSB repair in BALB/c-Trp53+/− cells, while downregulation was detected in C57BL/6-Trp53+/− cells. In fact, differences in DSB repair frequencies were mostly the result of decreases in homologous DSB repair in C57BL/6-Trp53+/− cells and/or increases in homologous repair in BALB/c-Trp53+/− cells. Notably, the increase in DSB repair is observed only when haploinsufficient for Trp53, which unmasks the lower fidelity repair in BALB/c.
Altogether, the siRNA screen identified 25 targets (Table 1) out of the 148 genes tested. The first-neighbor interactions among genes were mapped as shown in Figure 2. The targets gathered into two clusters indicating alterations in DNA replication (polymerases) and DSB repair (FA and BRCA) in BALB/c-Trp53+/− MEFs. These two clusters were connected by HR proteins and the RecQ helicase BLM. Strikingly, 12 of these 25 hit genes have been connected to crosslink DNA repair processes in the literature (Table 1).
Interactions among the gene targets identified in the siRNA screen. First neighbor interactions of gene targets showing differential repair efficiencies in MEFs derived from C57BL/6-Trp53+/− and BALB/c-Trp53+/− mice are shown. Rdm1 and Spo11 were omitted as these genes did not show connections with the network. Clusters of targets with common functions are highlighted (Polymerase, Helicase, FA, and BRCA clusters). The gene targets of siRNA that altered repair in C57BL/6-Trp53+/− MEFs, but not BALB/c-Trp53+/− MEFs, are shown in red. The relative effect is shown as a log2 ratio of repair efficiency for C57BL/6J-Trp53+/− and BALB/c-Trp53+/− MEFs ranging from 0.39–0.75 (1.3 to 1.7-fold) (Table 1).
DNA damage processing monitored by immunofluorescence microscopy
Radiation-induced DSBs can be repaired by different repair pathways,19 whereas DSBs generated during crosslink repair are subject to HR exclusively.20, 21 Having identified several crosslink repair proteins in the screen, we further dissected differences in the process of DNA damage removal after treatment with a crosslinking (mitomycin C, MMC) versus radiomimetic (bleomycin) drug. First, we visualized and quantified appearance and disappearance of DSBs by using antibodies against 53BP1, immunofluorescence microscopy and quantitative image analysis. Figure 3a shows that following MMC-treatment, BALB/c-Trp53+/− MEFs displayed a sharp increase in 53BP1 foci numbers. This rise was not observed with C57BL/6-Trp53+/− MEFs, even though the same cells showed 53BP1 foci accumulation after bleomycin incubation (data scattering for 53BP1 in bleomycin-treated BALB/c-Trp53+/− MEFs might be related to their radiosensitivity). To initiate HR, broken DNA ends need to be nucleolytically processed, followed by protection of single-strand DNA by RPA coverage.22 When we quantified RPA foci, we observed a significantly more pronounced accumulation 3–6 h after MMC incubation in BALB/c-Trp53+/− MEFs (Figure 3b). Replacement of RPA with Rad51 on single-stranded DNA during nucleofilament formation was monitored by Rad51 foci numbers, which were observed to be significantly higher in C57BL/6-Trp53+/− cells 1 h after MMC-treatment (Figure 3c). Accumulation of Rad51 foci was noticeable also in BALB/c-Trp53+/− MEFs but was delayed. Crosslink repair is triggered by stalled replication forks.21 Another mechanism causing replication fork arrest is pharmacological inhibition of poly(ADP-ribose)polymerase (PARP), presumably via the accumulation of unrepaired base damage at the fork, which is followed by conversion into DNA breaks and HR repair.23 After PARP inhibitor(NU1025) treatment, we observed a significant increase of 53BP1 foci in BALB/c-Trp53+/− but not C57BL/6-Trp53+/− MEFs (Figure 3d), suggesting that replication-associated repair of DSBs is compromised in general in BALB/c-Trp53+/− (not only after crosslinking). Comparative analysis of γH2AX foci indicated that after PARP inhibitor treatment foci accumulate in C57BL/6-Trp53+/− MEFs to a lesser extent than in BALB/c-Trp53+/− MEFs, qualitatively reflecting the corresponding 53BP1 foci result. The difference between NU1025-induced γH2AX foci in the two strains did not fully reach statistical significance (P=0.0571). This finding suggested that 53BP1 foci capture the critically different step, possibly DSB removal, during replication fork recovery better than γH2AX foci, which mark not only DSBs but also other DNA lesions, in particular stalled replication forks.24
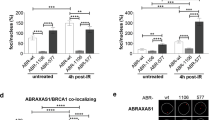
Immunofluorescence analysis of cells from Balb/c-Trp53+/− and C57BL/6-Trp53+/− mice. (a–c) MEFs were treated without (untreated) or with 2.6 μM MMC for 1 h or with 10 mU/ml bleomycin for 24 h, followed by reincubation in fresh medium. At the indicated incubation times post-treatment, cells were processed for immunolabeling of 53BP1 (a), RPA (b), or Rad51 (c). Representative images of 53BP1 immunostained and DAPI stained nuclei are depicted in (a). Immunolabeled foci were scored by automated quantification and mean numbers of foci per cell in the total cell population calculated from 4–8 slides with 25–45 (53BP1), 11–25 (RPA) and 10–31 (RAD51) nuclei each. Maximum mean scores were set to 100% for each experimental day and relative percentages calculated for each single value (corresponding to one slide). Hundred percent values represent the following average numbers of foci per cell: 53BP1: 9 (MMC and bleomycin); RPA: 1; Rad51: 3. Mean values (percentages) and s.e.m. values are shown graphically (*P<0.05; **P<0.01; ***P<0.001). (d) MEFs were incubated with 1 μM NU1025 for 24 h and immediately fixed for 53BP1 foci (left) or γH2AX (right) foci detection and quantification as in Figure 3a. Hundred percent represent 2 53BP1 and 15 γH2AX foci per cell, respectively. (e) MECs were incubated with 1 μM NU1025 for 24 h and 53BP1 foci quantified as MEFs in Figure 3a. Hundred percent represent 5 53BP1 foci per cell.
To validate our findings obtained with MEFs in mammary epithelial cells (MECs), that is, cells derived from the organ affected by breast cancer, we measured homologous DSB repair in MECs (Supplementary Figure 4) and performed immunofluorescence microscopic analysis following PARP inhibitor treatment of MECs exactly as in MEFs (Figure 3e). The results demonstrate that homologous DSB repair in MECs from BALB/c-Trp53+/− is 3.8-fold elevated compared with C57BL/6-Trp53+/− (P=0.0055). After PARP inhibitor treatment, we observed significant increases of 53BP1 foci both in BALB/c-Trp53+/− and C57BL/6-Trp53+/− MECs, however, to a significantly lesser extent in C57BL/6-Trp53+/− MECs compared with BALB/c-Trp53+/− MECs. Taken together, the results obtained with MECs confirmed our observations obtained with MEFs regarding a derepression of homologous DSB repair in BALB/c-Trp53+/− mice and exacerbated accumulation of DNA damage after replication arrest.
Functional analysis of the FA pathway
Crosslink repair dysfunction is a hallmark of FA patient cells, and this feature is made use of in diagnostic patient classification via chromosome breakage analysis following crosslinker-treatment.25 Cytogenetic analysis of MEFs at MMC concentrations of 0, 10, 50 and 100 ng/ml revealed a subtle increase of breakage events in BALB/c-Trp53+/− cells, as indicated by a lower percentage of BALB/c-Trp53+/− cells (66%+/−3) without any breaks after exposure to 10 ng/ml MMC compared with C57BL/6-Trp53+/− cells (75%+/−4), but mean values from three independent experiments did not fully reach statistical significance (P=0.0705, data not shown). Cell cycle analysis to detect a G2 arrest in response to crosslinker-treatment is another method, which more recently has successfully been applied to identify FA patients.26 The results with murine MEFs revealed that G2-phase accumulation was more pronounced in cells derived from BALB/c-Trp53+/− compared with C57BL/6-Trp53+/− mice without and after MMC-exposure (Figure 4). To exclude that the difference in G2-phase cells might simply reflect a difference in the rates of proliferation between the cell types, we determined BrdU incorporation in 3–4 independent experiments. The results showed that 57%+/−2 of BALB/c-Trp53+/− and 55%+/−1 of C57BL/6-Trp53+/− MEFs were BrdU-positive, without statistical significance of the 4% difference (P=0.1195). These results are consistent with a moderate FA pathway defect in BALB/c-Trp53+/−.
Functional analysis of the FA pathway. G2-phase accumulation. C57BL/6-Trp53+/− and BALB/c-Trp53+/− MEFs were treated without or with 10 ng/ml MMC. Forty-eight hours later the percentage of cells in the G2-phase of the cell cycle was determined by flow cytometric DNA content analysis after DAPI staining. Columns, mean values from three measurements each; bars, s.d. Note that C57BL/6-Trp53+/− cells contained ∼30% tetraploid cells, which were excluded from the calculation of cellular G2-phase fractions.
FA pathway components are subdivided into proteins involved in FancD2 and FancI monoubiquitylation and others acting downstream of that step, which more directly participate in HR repair.20 To discriminate between these two possibilities, we performed immunoblotting specific for FancD2. We noticed the appearance of a more slowly migrating band, indicative of FancD2 ubiquitylation, after bleomycin treatment both in BALB/c-Trp53+/− and C57BL/6-Trp53+/− MEFs (Figure 5a). From this result, the two FA genes FancE and FancB, which were identified in the screen (Figure 1c, Table 1) and which are positioned upstream of FancD2 in the FA pathway, were unlikely to be altered in BALB/c-Trp53+/− mice. However, the screening hit Palb2 is positioned downstream of FancD2. To determine whether a genetic change in Palb2 was causal to homologous repair deregulation in BALB/c-Trp53+/−, we co-transfected MEFs with plasmid for expression of exogenous murine Palb2, substrate Δ-EGFP/3′EGFP and I-SceI expression plasmid (Figure 5b). Palb2 expression in C57BL/6-Trp53+/− MEFs enhanced homologous repair 2.6-fold (P=0.0313), whereas the 1.6-fold increase in BALB/c-Trp53+/− MEFs was not statistically significant (P=0.1563). For comparison, we assessed the effect of expressing exogenous PALB2 in PALB2-deficient versus-proficient cells, namely in the PALB2-mutated patient fibroblast cell line EUFA134127 and the human fibrosarcoma cell line HT1080. Expression of human PALB2 in EUFA1341 cells raised the homologous repair frequency 4.7-fold (P=0.0039); the 3.2-fold rise in HT1080 cells was not statistically significant. The pronounced increase of homologous DSB repair in EUFA1341 but not BALB/c-Trp53+/− MEFs made it unlikely that lack of Palb2 functionality was causal to the homologous DSB repair phenotype in these MEFs. To understand whether downstream of Palb2, expression of the FANCD1/BRCA2 gene was differentially affected in the two MEF species, we performed immunoblotting and qRT–PCR experiments. Figure 6a shows that equivalent levels of homologous repair and signaling proteins like ATM, MRE11, RAD50, BLM, Palb2, or Rad51 were detected. Different from the other MRE11-RAD50-Nibrin complex components, Nibrin levels were 1.7-fold elevated in BALB/c-Trp53+/− MEFs, which might be explained by genotoxic stress due to a defect in replication-associated DNA repair in these cells (Figure 3d).28 Constitutive deregulation of Nibrin expression in BALB/c-Trp53+/− MEFs is unlikely, because it should correspondingly affect the levels of the other complex components MRE11 and RAD50.28 Most strikingly, BRCA2 protein expression in BALB/c-Trp53+/− MEFs was less than half of the level in C57BL/6-Trp53+/− MEFs. mRNA levels of BRCA2 were similar in both cell types (Figure 6b), suggesting that post-transcriptional processes differentially affected BRCA2 protein levels. To understand whether reduced BRCA2 protein contributes to the functional change in replication-associated HR in BALB/c-Trp53+/−, we analyzed NU1025-induced 53BP1 foci formation following BRCA2 silencing in MEFs from the two Trp53+/−strains. The results are displayed in Figure 6c and show that after BRCA2 knockdown a statistically significant difference between 53BP1 foci numbers in BALB/c-Trp53+/− and C57BL/6-Trp53+/− MEFs was indeed no longer observed (P=0.1143).
Dissection of the FA pathway. (a) FancD2 protein analysis. MEFs from BALB/c-Trp53+/− and C57BL/6-Trp53+/− mice were treated with H2O or with 10 mU/ml bleomycin for 24 h followed by protein extraction and western analysis of 110 μg of total protein and immunodetection with antibodies directed against the DSB repair protein FancD2. Loading of the corresponding lanes is visualized by Ponceau staining of total proteins. An additional more slowly migrating FancD2 band (asterisk) was detected particularly in drug-treated samples and indicates mono-ubiquitination of FancD2. (b) Homologous DSB repair activities in response to transfection with Palb2 expression plasmid. C57BL/6-Trp53+/− and BALB/c-Trp53+/− MEFs were transfected with homologous DSB repair substrate (Δ-EGFP/3′EGFP), pCMV-I-SceI and expression plasmid for murine Palb2 (DIRES-Palb2) or empty vector (DIRES-DGFPII). EGFP-positive cells were quantified 48 h after transfection and normalized for transfection efficiencies. Palb2-mutated EUFA1341 human fibroblast and HT1080 human fibrosarcoma cells were correspondingly analyzed by use of the expression plasmid for human PALB2 (POZ-PALB2) or empty vector (POZ). Mean DSB repair frequencies and s.e.m. values are shown (n=6–9). Frequencies in cells transfected with empty vector were defined as 100%. Asterisks indicate a statistically significant difference (Wilcoxon-matched pairs test, two-tailed, C57BL/6-Trp53+/−: P=0.0313; EUFA1341: P=0.0039; *P<0.05; **P<0.01). Levels of total Palb2 in C57BL/6-Trp53+/− and BALB/c-Trp53+/− cells were visualized by immunoblotting with Palb2-specific antibodies, exogenous PALB2 in EUFA1341 and HT1080 cells by antibodies directed against the HA-tag. α-tubulin served as loading control. Framed images were derived from the same western blot and autoradiographic exposure. An arrow marks the band specific for HA-tagged PALB2 in human cells.
Expression analysis of DSB repair factors. (a) Comparative analysis of DSB repair protein levels. Endogenous levels of DSB repair proteins in MEFs from C57BL/6-Trp53+/− and BALB/c-Trp53+/− mice were visualized after electrophoresis of extracts containing 60 μg of total protein on 12% SDS–PAGE or NuPAGE Novex 4–12% gradient gels and immunblotting with antibodies directed against the indicated proteins including the loading controls α-tubulin and TATA-binding protein (TBP). Framed images were derived from the same western blot and autoradiographic exposure. In the comparative graphical presentation of DSB repair, protein levels columns indicate relative band intensities quantified from 2–4 independent immunoblots after normalization for protein loading each. Values for C57BL/6-Trp53+/− were set to 100% for each immunodetection. Columns indicate mean values; bars indicate s.d. (b) Quantitative BRCA2 mRNA expression analysis by RT–PCR. C57BL/6-Trp53+/− and BALB/c-Trp53+/− MEFs were either left untreated or transfected with a DNA and siRNA mixture as described in the legend to Figure 1 including either non-silencing siRNA control siRNA or pools of four siRNAs directed against BRCA2. After 24 h, RNA was extracted, cDNAs synthesized and the mRNA expression levels of the BRCA2 gene determined by RT–PCR. Mean expression levels in untreated and non-silencing siRNA-transfected C57BL/6-Trp53+/− MEFs, respectively, were set to 1.0 and relative DNA levels calculated from a standard curve. Mean values and s.e.m. were obtained from six independent measurements. *P<0.05; (c) Immunofluorescence analysis of Balb/c-Trp53+/− and C57BL/6-Trp53+/− MEFs after BRCA2 knockdown. Low passage BALB/c-Trp53+/− or C57BL/6-Trp53+/− MEFs were transfected with a pool of four different siRNAs directed against BRCA2, cultivated for 24 h, then, treated with 1 μM NU1025 for 24 h and immediately fixed for 53BP1 foci detection and quantification in 53BP1 foci-positive cells. Mean values (percentages) and s.e.m. values for four slides each are shown (*P<0.05). Hundred percent represent 18 53BP1 foci.
Discussion
The striking molecular overlap between FA and breast cancer susceptibility (BRCA) is exemplified best by the existence of at least four genes (BRCA2/FANCD1, BRIP1/FANCJ, PALB2/FANCN, RAD51C/FANCO), which concomitantly represent FA and BRCA pathway genes.26, 29, 30 Moreover, both FA and BRCA pathway genes have a role in homologous repair.20, 26, 29, 30 In this study, we performed a siRNA screen to identify genes that differentially affect homologous repair in cells from BALB/c-Trp53+/− that are susceptible versus C57BL/6-Trp53+/− mice that are resistant to mammary tumors. Screening relied on repair measurements with an EGFP-based DNA substrate that was recently demonstrated to detect a repair increase in individuals with hereditary breast cancer risk.16 The same substrate was also shown to detect HR-regulatory functions of p53 particularly well, because homologies within the two-mutated EGFP genes are <200 bp, a critical limit for HR fidelity control.31 Our siRNA screen led to the identification of 25 genes, among which we identified partially overlapping groups of genes that had previously been linked with crosslink repair (11 genes), FA and BRCA1/2 (8 genes), translesion synthesis(six genes) and repair replication (five genes) (Table 1). The significance is supported by the fact that these groups are functionally connected and that the hits form a network of physical and functional interactions.
We did not identify even a single-gene belonging to the canonical NHEJ pathway. In particular, we did not identify DNA-PKcs, even though downregulation of protein expression was verified (data not shown). Moreover, treatment of MEFs with DNA-PK inhibitor did not alter homologous repair frequencies, but the ATM/ATR inhibitor caffeine, which exerts control on homologous repair factors,19 exerted significantly different effects on BALB/c-Trp53+/− and C57BL/6-Trp53+/− MEFs. Therefore, we consider it unlikely that the reduced DNA-PKcs levels in BALB/c can explain the difference between homologous repair activities of the two strains. Measurements of microhomology-mediated NHEJ further excluded a major difference in the alternative NHEJ pathway. In conclusion, our results from DSB repair measurements and siRNA screening uncovered a difference in the homologous and crosslink repair pathways.
Crosslink repair can roughly be subdivided into the following sequentially occurring repair stages: replication fork pausing and damage signaling for activation of downstream repair and checkpoint proteins, incisions causing one-ended DSB formation, translesion synthesis, end resection and adduct removal, and Rad51-dependent HR.20, 21 Here, we noticed accumulation of 53BP1 foci immediately after crosslinker-treatment in BALB/c-Trp53+/− MEFs indicating accumulation of DSBs. We interpreted this result such that after fork stalling due to the crosslink, the initial incision step next to the crosslink was active in BALB/c-Trp53+/− MEFs, but removal of the resulting DSBs was delayed.20, 21 Further supporting proficiency of endonucleolytic cleavage, we also failed to detect reduced FancD2 ubiquitylation, which occurs after cleavage and before end processing.20 From this, we concluded that the crosslink repair pathway was compromised downstream of the incision step in BALB/c-Trp53+/− MEFs. Single-strand DNA end formation was monitored by RPA foci analysis, which revealed accumulation of processed ends in BALB/c-Trp53+/− MEFs. These findings indicated that nucleolytic processing itself was not compromised. However, further downstream Rad51 nucleoprotein filament formation was delayed and less productive, which could explain increased RPA foci numbers. Thus, the critical step discriminating the response of the different types of MEFs could either be adduct removal or Rad51 loading onto the DNA overhang. From the fact that we also observed 53BP1 accumulation in BALB/c-Trp53+/− MEFs and MECs after PARP inhibitor treatment, which causes replication fork stalling independently of crosslinks,23 adduct removal by nucleotide excision repair proteins can be excluded, thus, leaving the possibility of deficient Rad51 filament formation in BALB/c-Trp53+/− MEFs.
Hits from our siRNA screen, which promote Rad51 filament formation and, therefore, were the best candidates for a BALB/c-specific genetic alteration, were Palb2, Xrcc2, Xrcc3 and possibly the BRCA2-associated proteins Hmg20b and Rdm1, the latter one showing sequence similarity to Rad52 and involvement in the cellular cisplatin response.20, 32, 33 Xrcc2 and Xrcc3 polymorphisms have been associated with breast cancer risk.34, 35 Most recently, exome sequencing implicated a truncating Xrcc2 mutation in the pathogenesis of FA.36 However, among the five above-mentioned genes PALB2/FancN has most convincingly been defined as both BRCA and FA gene.26, 29 In support of subtle FA pathway dysfunction, BALB/c-Trp53+/− MEFs showed signs of elevated chromosome breakage and G2-phase accumulation after MMC-exposure, that is, manifestations resembling hallmarks for FA patient diagnosis.
After silencing we would expect to see an activity decrease, if the silenced gene is required, but only a small effect, when an endogenous defect already affects the same gene or pathway. Focusing on corresponding screening hits for which we noticed >20% reduction in C57BL/6-Trp53+/-, but <10% frequency change in BALB/c-Trp53+/− MEFs, identified Palb2, FancE, RecQL, Acaca, Polymerase(Pol)ɛ3, Hmg20b and FancB, that is, enriched FA/BRCA pathway components (Palb2, FancE, FancE) and associated factors (Acaca, Hmg20b)(Figure 1c, Table 1). Among those, we assigned the biggest DSB repair difference and the lowest P-value to Palb2. However, when we tried to rescue a potential Palb2 defect in BALB/c-Trp53+/− MEFs by expressing wild-type protein, we observed a smaller effect on homologous repair compared with C57BL/6-Trp53+/− MEFs. This result could be explained by a defect downstream of Palb2, and in agreement with this idea we observed significantly reduced BRCA2 protein levels in BALB/c-Trp53+/−. Consistently, PARP inhibitor-induced 53BP1 foci formation after BRCA2 silencing in MEFs from the two Trp53+/-strains indicated an effect of reduced BRCA2 protein levels on replication-associated HR in BALB/c-Trp53+/−. The fact that BRCA2 was not identified in the screen could be due to the fact that it is an essential gene,37 and indeed, we frequently observed cytotoxicities 48 h after BRCA2-siRNA/plasmid DNA co-transfection. Comparison of qRT–PCR and immunoblot data indicated BRCA2 regulation at the post-transcriptional level. BRCA2 is known to be post-translationally modified38 and complex formation with other proteins such as with Palb2 prevent destabilization.39, 40 Therefore, numerous possible explanations for reduced BRCA2 levels in BALB/c-Trp53+/− remain and have to await further investigations.
FA proteins, including BRCA2/FancD1, act on replication restart at stalled forks.20, 41 Consistent with defective HR-dependent recovery of broken forks in BALB/c-Trp53+/− cells, we noticed sharp accumulation of DSBs after PARP inhibitor treatment, a slight rise of cells in G2 even without treatment, and increased Nibrin protein levels. Partially compromised BRCA2 functions in combination with knockdown of one of the other hits of our screen involved in the bypass of lesions (translesion synthesis polymerases) or resolution of structural obstacles (BLM) may exacerbate replication stress in BALB/c-Trp53+/− cells, thereby triggering ATM/ATR-mediated activation of the homologous repair machinery.20 Indeed, we observed an increase of homologous repair in BALB/c-Trp53+/− but not C57BL/6-Trp53+/− cells after knockdown of Polζ subunit Rev7 (Mad2l2), Polκ, Polη, Polɛ or BLM. The same translesion synthesis polymerases have also been linked with crosslink repair.21, 42 Moreover, polymerases and cofactors identified in our siRNA screen, such as Polɛ, its binding partner Paxip1, and Polδ-binding protein Trex2 have a role in heteroduplex DNA extension during HR.43, 44, 45 Interestingly, genetic variation in genes encoding Polζ subunits associates with breast cancer risk.46
Elevated basal levels of replication stress may also explain why we measured increased homologous repair in BALB/c-Trp53+/− MEFs and MECs even without knockdown (Figure 1a and Supplementary Figure 4). In our experiments, Rad51 was not loaded onto single-strand DNA in BALB/c-Trp53+/− as quickly as in C57BL/6-Trp53+/− MEFs, therefore, broken DNA ends were less protected against nucleolytic processing, which is in agreement with the observed accumulation of RPA foci after MMC-treatment. Stalled replication fork degradation by MRE11 was reported to be the consequence of loss of BRCA2 function,41 which is compatible with reduced BRCA2 levels in BALB/c-Trp53+/− MEFs. Extensive end processing promotes a shift from high-fidelity HR to error-prone homologous repair processes,47 which are major sources for genomic rearrangements, such as SSA between repeat sequences or long-tract gene conversion events associated with LOH.48 Consistently, when using the very same homologous repair substrate as applied to our screen in this study on cells from breast cancer predisposed individuals carrying mutations in BRCA2, we measured a two- to threefold increase.49 Therefore, we propose that increased homologous DSB repair in BALB/c-Trp53+/− MEFs after knockdown of many of the hit genes including Palb2 was because of further stimulation of aberrant DSB repair due to additional deficiencies in the FA/BRCA pathway and/or due to additional replication stress signaling. Notably, Sy et al.39 also observed HR increase rather than decrease after MRG15 knockdown causing reduced Palb2 and BRCA2 levels. These and further observations by other groups suggest that HR needs to be tightly controlled in terms of the BRCA2/Rad51 balance, to avoid both excessive and unscheduled HR events, as well as alternative error-prone homologous exchange mechanisms for maintenance of genomic stability.22
Even though BALB/c mice carry a defect in the FA/BRCA pathway, additional loss of control by Trp53-heterozygosity appears to be necessary for deregulated homologous repair, as repair frequencies were indistinguishable between cells from Trp53+/+ mice of the two strains (not shown). p53 was previously shown to control Rad51-dependent HR6, 7 and to have a role in crosslink repair.50, 51 Therefore, it is conceivable that Trp53-heterozygosity combined with a subtle crosslink repair defect sums up to a significant deficiency in DSB repair. By extension, it is conceivable that reduced levels of the BRCA gene products p53 and BRCA2, or the absence of another factor X in the FA/BRCA pathway in combination with variants of modifier genes including DMBT152 reduce latency of mammary tumorigenesis in BALB/c-Trp53+/− mice. Complex tumorigenesis models integrating combined effects of low-penetrance modifier alleles and epigenetic changes, have led to a paradigm shift from single-gene genotyping to more comprehensive diagnostic tools such as functional analysis in biomarker research.53, 54 Thus, BALB/c-Trp53+/− mice not only represent a LFS, but additionally a complex breast cancer mouse model inspiring future studies for the assessment of the effects of various combined subtle defects in the DSB and crosslink repair family of genes on breast cancer predisposition and therapeutic responses.
Materials and methods
Mouse strains and cell culture
BALB/c-Trp53+/− mice were generated as described previously55 by backcrossing (C57BL/6 × 129/Sv) Trp53−/− mice onto the BALB/cMed-strain for >13 generations. MEFs were isolated from individual embryos at day 13.5 to allow confirmation of genotypes. Collagenase was used to obtain fibroblasts, which were maintained in alpha MEM supplemented with 10% FBS and β-mercaptoethanol until confluent. MEFs were frozen in pooled aliquots so that assays could be performed with cells at passage ⩽3. EUFA1341 human fibroblasts27 and HT1080 human fibrosarcoma cells were cultivated in high-glucose DMEM supplemented with 15% FBS and 2 mM L-glutamine. MECs from three mice were pooled for each set of experiments and cultivated on plates coated with Collagen I (ScienCell Research Laboratories, Carlsbad, CA, USA) and maintained in EpiCult-B medium (StemCell Technologies, Sirocco, France) containing 10 ng/ml mEGFP (Sigma, Deisenhofen, Germany), 10 ng/ml human basic fibroblastic growth factor (Sigma), 4 μg/ml Heparin (Sigma), 5 μg/ml Insulin (Sigma), 1 μg/ml Hydrocortisone (Sigma) and 10% FBS up to passage 3. Cells were routinely tested negative for Mycoplasma contamination.
DSB repair assay and plasmid construction
Cells were transfected with a mixture containing FugeneHD (Roche, Penzberg, Germany) and either repair plasmid Δ-EGFP/3′EGFP or EJ-EGFP (Figure 1a),31 pCMV-I-SceI (I-SceI expression), and filler plasmid pBS. For expression of human PALB2, we additionally included POZ-PALB2.27 The murine Palb2 expression construct DIRES-Palb2 was prepared by PCR amplification of overlapping fragments of the Palb2 mRNA (nucleotides 28–1671 and 1331–3720) from mammary glands of C57BL/6 mice. The complete open reading frame was cloned into the pIRES-hrGFP II vector (Agilent Technologies, Santa Clara, CA, USA). The IRES-GFP sequences were deleted in the construct used for DSB repair assays.
For assessment of transfection efficiency pBS was replaced with wtEGFP expression vector (based on Δ-EGFP/3′EGFP backbone) in split samples. Twenty-four hours post-transfection, the percentage of green fluorescent cells was assessed on a FACSCalibur (BD Biosciences, Heidelberg, Germany) with 488 nm laser excitation and detection of gated live cells (SSC/FSC dot plot) in the FL1/FL2 channels (Figure 1b).31 To verify that the results obtained upon split sample transfection are comparable with the ones for co-transfection with a fluorescing reporter plasmid, we measured homologous DSB repair in MEFs under both conditions (Supplementary Figure 1A). Split sample transfection with wtEGFP expression vector confirmed the twofold increase in the homologous DSB repair frequency (Δ-EGFP/3′EGFP) in BALB/c-Trp53+/− versus C57BL/6-Trp53+/− depicted in Figure 1a. Measurements of homologous DSB repair frequencies upon mCherry expression vector (pmCherry-N1 from Clontech, Heidelberg, Germany) co-transfection confirmed more active homologous DSB repair in BALB/c-Trp53+/− MEFs. However, frequencies normalized for mCherry were lower for both strains thereby reaching the detection limit with MEFs from C57BL/6-Trp53+/−.
The following drugs were added 1 h pretransfection: KU-55933 (ATM, KuDOS Pharmaceuticals, Cambridge, UK), NU7441 (DNA-PK, KuDOS) and caffeine (ATM/ATR, Sigma-Aldrich, Deisenhofen, Germany). Transfection efficiencies in a typical experiment as depicted in Supplementary Figure 1B varied between triplicate samples with a s.d. of 3–8% for DMSO-treated and 13–18% for caffeine-treated cells for the two MEF types (relative to mean values set to 100%).
DSB repair analysis under conditions of RNA interference
Passage 2 MEFs were plated at a density of 100 000 cells per six-well. After 24 h, cells were transfected with a mixture containing Hiperfect (Qiagen, Hilden, Germany), 24 pmol siRNA (Qiagen) and 1.25 μg plasmid DNA (Δ-EGFP/3′EGFP,31 pCMV-I-SceI, pBS). To determine transfection efficiencies, pBS was replaced with wtEGFP plasmid. 48 h post-transfection cells were analyzed for EGFP-positivity by flow cytometry. siRNAs were used as pools of four target gene-specific siRNA duplexes (HP GenomeWide siRNA design by Qiagen, potent knockdown guaranteed for ⩾2 individual siRNAs) during all screening rounds. Single siRNA duplex was used only in control experiments in Supplementary Figures 2A and 3. Non-silencing siRNAs served as negative controls (AS). GFP-specific siRNA-silenced GFP expression down to 1%.
Screening procedure and data evaluation
The screen was performed twice for each target-specific siRNA pool (Supplementary Table 1 for genes, log2 ratios and P-values; Supplementary Table 2 for siRNA sequences) with four independent transfections for repair and two transfections for wtEGFP controls each. For calculation of DSB repair frequency, the percentage of fluorescent cells in the individual sample was corrected for the transfection efficiency (wtEGFP samples) and then normalized to the mean frequency obtained in parallel samples with non-silencing siRNA. Statistical significance was calculated with non-parametric Mann–Whitney U-test (GraphPad Software, Inc., San Diego, CA, USA). To score differences in BALB/c-Trp53+/− versus C57BL/6-Trp53+/− MEFs, normalized repair frequencies were subjected to logarithmic transformation to the base two and inserted in the formula: ratio of DSB repair frequency difference=log2(normalized DSB repair frequency [BALB/c-Trp53+/-])—log2(normalized DSB repair frequency [C57BL/6-Trp53+/−]).
Integration of efficiency of repair with protein interactions
The human protein interactome was built with data downloaded from the NCBI database 13, February 13 (ftp://ftp.ncbi.nih.gov/gene/GeneRIF). Interactions were formatted in the Cytoscape readable simple interaction format and loaded into Cytoscape.56 For the siRNA targets that showed differences in repair activities in BALB/c-Trp53+/− versus C57BL/6-Trp53+/− MEFs (Table 1, P<0.01), the mouse Gene IDs were converted from Mouse Entrez Gene IDs to Human Entrez Gene IDs (HomoloGene Build 65; ftp://ftp.ncbi.nih.gov/pub/HomoloGene/build65/) and mapped onto the protein interactome. The log2 gene expression ratio between the two strains (BALB/c:C57BL/6) was overlayed onto the interactome to integrate expression data generated in the current study with protein interaction data from the NCBI database.
Trp53 LOH analysis
To verify maintenance of the heterozygous Trp53 genotype, genomic DNA was extracted from MEFs grown in parallel to cells used in the screen (Figure 1b) following the Fermentas Molecular Biology Tools protocol. Genomic DNA was used in multiplex PCR specific for the wild-type or mutant Trp53 locus.55
Immunofluorescence analysis
MEFs were incubated with bleomycin (10 mU) for 24 h, MMC (2.6 μM) for 45 min, or MEFs and MECs with NU1025 (1 μM) for 24 h (Sigma-Aldrich). Thereafter, cells were either fixed immediately with 3.7% formaldehyde in phosphate-buffered saline and permeabilized with 0.5% TritonX-100, or further incubated without drugs. Primary antibodies used were polyclonal antibodies against 53BP1 (NB100-304, Novus Biologicals, Littleton, CO, USA), Rad51 (H-92, Santa Cruz Biotechnology, Heidelberg, Germany), and monoclonal against RPA (RPA34-19, Calbiochem, Darmstadt, Germany), γH2AX (Clone JBW 301, Millipore, Eschborn, Germany). Secondary antibodies were AlexaFluor555 labeled (Invitrogen, Karlsruhe, Germany). Images were acquired on an Olympus fluorescence microscope BX51 with × 100 oil immersion objective equipped with cooled charge-coupled device (CCD) camera (Colorview 12, Olympus, Tokyo, Japan). Foci quantification was carried out with Cell^F imaging software version 2.5 (Olympus Soft Imaging Solutions, Münster, Germany). Threshold and minimal focus size were maintained throughout one set of simultaneously treated and processed samples.
Western blotting
Cells were treated with bleomycin (5 mU/ml) for 24 h, with MMC (2.6 μM) for 45 min or solvent as indicated. For knockdown verification under screening conditions, cells were harvested 48 h post-transfection. Cell pellets were extracted, separated electrophoretically (SDS–PAGE or NuPAGE-Novex-gradient-gels, Invitrogen), and transferred to polyvinylidene difluoride (PVDF) membrane.49 Proteins were detected by incubation with murine antibodies against ATM (mAb 5C2, Abcam, Cambridge, UK), DNA-PKcs (ab-4, Neomarkers, Fremont, CA, USA), RAD50 (mAb GTX70228, GeneTex, Irvine, CA, USA), TATA-binding protein (TBP) (mAb 1TBP18, Abcam), α-tubulin (mAb DM1A, Abcam), rabbit antibodies against ATR (polyclonal Ab-2, Calbiochem), BLM (polyclonal ab476, Abcam), FancD2 (polyclonal ab2187, Abcam), MRE11 (polyclonal NB100-142, Novus Biologicals), Nibrin (mAb Y112, Epitomics), PALB2 (polyclonal BCN4785, Antibody BCN), Rad51 (polyclonal, H-92, Santa Cruz Biotechnology), rat antibodies against HA-tag (mAb 3F10, Roche) or goat antibodies against BRCA2 (polyclonal N-19, Santa Cruz) and visualized by chemiluminescence (SuperSignal West Dura Extended Duration Substrate, Pierce, Rockford, IL, USA). Densitometric quantification was carried out with AlphaView software (Cell Biosciences, Heidelberg, VIC, Australia).
Quantitative real-time PCR
In order to confirm knockdown of target genes by co-transfection with siRNA, qRT–PCR was performed 24–48 h post-transfection. Total RNA was extracted with RNeasy-Mini-Plus-Kit and reverse transcription carried out with QuantiTect-Reverse-Transcription-Kit (Qiagen). qRT–PCR on cDNA samples was carried out in triplicates on an ABI-Prism-7700-Gene-Analyzer (Applied Biosystems, Carlsbad, CA, USA) using QuantiTect-SYBR-Green-PCR-Kit and gene-specific reverse and forward primers (QuantiTect-Primer-Assay). We examined mRNA downregulation for the 25 hit genes from Figure 1c and Table 1, but failed to confirm knockdown for Usp1, Hmg20B and FancB, likely due to accelerated cell death that we reproducibly observed in these knockdown cultures (data not shown). For the hit genes Xrcc2, Palb2 and Acaca from the FA/BRCA cluster and for BRCA2, we calculated cDNA levels relative to a standard curve as displayed in Supplementary Figure 2B and Figure 6b.
Chromosome breakage, cell cycle analysis
MEFs from BALB/c-Trp53+/− and C57BL/6-Trp53+/− mice were grown in Amniopan (PAN Biotech, Aidenbach, Germany) and exposed to MMC at final concentrations of 0, 10, 50 or 100 ng/ml for 36 h. Metaphase preparation followed standard procedures. A minimum of 50 complete metaphases from Giemsa-stained slides for either mouse strain and each MMC concentration were scored regarding chromosome breakage rates.26
Parallel cultures of MEFs were left untreated or exposed to 10 ng/ml MMC for 48 h, harvested and stained in medium containing 15 μg/ml Hoechst dye 33342 (Molecular Probes, Invitrogen). We gated on vital cells via propidium iodide exclusion. Split samples were stained with 1 μg/ml 4′,6-diamidino-2-phenylindole (DAPI) (Molecular Probes), univariate flow histograms recorded on an LSRII flow cytometer (Becton Dickinson, Heidelberg, Germany), and cell cycle distributions quantified using the MPLUS AV software package (Phoenix Flow Systems, San Diego, CA, USA).26 For assessment of the S-phase index, BALB/c-Trp53+/− and C57BL/6-Trp53+/− MEFs were labeled 1 h with BrdU by use of the FITC BrdU Flow Kit (BD Pharmingen, Heidelberg, Germany) according to the instructions of the manufacturer.
References
Tainsky MA, Bischoff FZ, Strong LC . Genomic instability due to germline p53 mutations drives preneoplastic progression toward cancer in human cells. Cancer Metastasis Rev 1995; 14: 43–48.
Shlien A, Tabori U, Marshall CR, Pienkowska M, Feuk L, Novokmet A et al. Excessive genomic DNA copy number variation in the Li-Fraumeni cancer predisposition syndrome. Proc Natl Acad Sci USA 2008; 105: 11264–11269.
Zhao X, Weir BA, LaFramboise T, Lin M, Beroukhim R, Garraway L et al. Homozygous deletions and chromosome amplifications in human lung carcinomas revealed by single nucleotide polymorphism array analysis. Cancer Res 2005; 65: 5561–5570.
Shao C, Deng L, Henegariu O, Liang L, Stambrook PJ, Tischfield JA . Chromosome instability contributes to loss of heterozygosity in mice lacking p53. Proc Natl Acad Sci USA 2000; 97: 7405–7410.
Lu X, Lozano G, Donehower LA . Activities of wildtype and mutant p53 in suppression of homologous recombination as measured by a retroviral vector system. Mutat Res 2003; 522: 69–83.
Bertrand P, Saintigny Y, Lopez BS . p53′s double life: transactivation-independent repression of homologous recombination. Trends Genet 2004; 20: 235–243.
Gatz SA, Wiesmüller L . p53 in recombination and repair. Cell Death Differ 2006; 13: 1003–1006.
Chompret A, Brugieres L, Ronsin M, Gardes M, Dessarps-Freichey F, Abel A et al. p53 germline mutations in childhood cancers and cancer risk for carrier individuals. Br J Cancer 2000; 82: 1932–1937.
Kuperwasser C, Hurlbut GD, Kittrell FS, Dickinson ES, Laucirica R, Medina D et al. Development of spontaneous mammary tumors in BALB/c p53 heterozygous mice. A model for Li-Fraumeni syndrome. Am J Pathol 2000; 157: 2151–2159.
Blackburn AC, Brown JS, Naber SP, Otis CN, Wood JT, Jerry DJ . BALB/c alleles for prkdc and cdkn2a interact to modify tumor susceptibility in trp53+/− mice. Cancer Res 2003; 63: 2364–2368.
Blackburn AC, McLary SC, Naeem R, Luszcz J, Stockton DW, Donehower LA et al. Loss of heterozygosity occurs via mitotic recombination in Trp53+/− mice and associates with mammary tumor susceptibility of the BALB/c strain. Cancer Res 2004; 64: 5140–5147.
Blackburn AC, Hill LZ, Roberts AL, Wang J, Aud D, Jung J et al. Genetic mapping in mice identifies DMBT1 as a candidate modifier of mammary tumors and breast cancer risk. Am J Pathol 2007; 170: 2030–2041.
Koch JG, Gu X, Han Y, El-Naggar AK, Olson MV, Medina D et al. Mammary tumor modifiers in BALB/cJ mice heterozygous for p53. Mamm Genome 2007; 18: 300–309.
Szpirer C, Szpirer J . Mammary cancer susceptibility: human genes and rodent models. Mamm Genome 2007; 18: 817–831.
Forbes SA, Bindal N, Bamford S, Cole C, Kok CY, Beare D et al. COSMIC: mining complete cancer genomes in the catalogue of somatic mutations in cancer. Nucleic Acids Res 2011; 39 (Database issue): D945–D950.
Keimling M, Deniz M, Varga D, Stahl A, Schrezenmeier H, Kreienberg R et al. The power of DNA double-strand break (DSB) repair testing to predict breast cancer susceptibility. FASEB J 2012; 26: 2094–2104.
Okayasu R, Suetomi K, Yu Y, Silver A, Bedford JS, Cox R et al. A deficiency in DNA repair and DNA-PKcs expression in the radiosensitive BALB/c mouse. Cancer Res 2000; 60: 4342–4345.
Mitchell J, Smith GCM, Curtin NJ . Poly(ADP-Ribose) polymerase-1 and dna-dependent protein kinase have equivalent roles in double strand break repair following ionizing radiation. Int J Radiat Oncol Biol Phys 2009; 75: 1520–1527.
Asaad NA, Zeng ZC, Guan J, Thacker J, Iliakis G . Homologous recombination as a potential target for caffeine radiosensitization in mammalian cells: reduced caffeine radiosensitization in XRCC2 and XRCC3 mutants. Oncogene 2000; 19: 5788–5800.
Thompson LH, Hinz JM . Cellular and molecular consequences of defective Fanconi anemia proteins in replication-coupled DNA repair: mechanistic insights. Mutat Res 2009; 668: 54–72.
Ho TV, Schärer OD, Translesion DNA . Synthesis polymerases in DNA crosslink repair. Environ Mol Mutagen 2010; 51: 552–566.
Schild D, Wiese C . Overexpression of RAD51 suppresses recombination defects: a possible mechanism to reverse genomic instability. Nucleic Acids Res 2010; 38: 1061–1070.
Heacock ML, Stefanick DF, Horton JK, Wilson SH . Alkylation DNA damage in combination with PARP inhibition results in formation of S-phase-dependent double-strand breaks. DNA Repair 2010; 9: 929–936.
Gagou ME, Zuazua-Villar P, Meuth M . Enhanced H2AX phosphorylation, DNA replication fork arrest, and cell death in the absence of Chk1. Mol Biol Cell 2010; 21: 739–752.
Auerbach AD, Rogatko A, Schroeder-Kurth TM . International Fanconi Anemia Registry: relation of clinical symptoms to diepoxybutane sensitivity. Blood 1989; 73: 391–396.
Vaz F, Hanenberg H, Schuster B, Barker K, Wiek C, Erven V et al. Mutation of the RAD51C gene in a Fanconi anemia-like disorder. Nat Genet 2010; 42: 406–411.
Xia B, Dorsman JC, Ameziane N, de Vries Y, Rooimans MA, Sheng Q et al. Fanconi anemia is associated with a defect in the BRCA2 partner PALB2. Nat Genet 2007; 39: 159–161.
Kavitha CV, Choudharty B, Raghavan SC, Muniyappa K . Differential regulation of MRN (Mre11-Rad50-Nbs1) complex subunits and telomerase activity in cancer cells. Biochem Biophys Res Commun 2010; 399: 575–580.
Walsh T, King MC . Ten genes for inherited breast cancer. Cancer Cell 2007; 11: 103–105.
Meindl A, Hellebrand H, Wiek C, Erven V, Wappenschmidt B, Niederacher D et al. Germline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility gene. Nat Genet 2010; 42: 410–414.
Akyüz N, Boehden GS, Süsse S, Rimek A, Preuss U, Scheidtmann KH et al. DNA substrate dependence of p53-mediated regulation of double-strand break repair. Mol Cell Biol 2002; 22: 6306–6317.
Marmorstein LY, Kinev AV, Chan GK, Bochar DA, Beniya H, Epstein JA et al. A human BRCA2 complex containing a structural DNA binding component influences cell cycle progression. Cell 2001; 104: 247–257.
Hamimes S, Arakawa H, Stasiak AZ, Kierzek AM, Hirano S, Yang YG et al. RDM1, a novel RNA recognition motif (RRM)-containing protein involved in the cell response to cisplatin in vertebrates. J Biol Chem 2005; 280: 9225–9235.
Lin WY, Camp NJ, Cannon-Albright LA, Allen-Brady K, Balasubramanian S, Reed MW et al. A role for XRCC2 gene polymorphisms in breast cancer risk and survival. J Med Genet 2011; 48: 477–484.
He XF, Wei W, Su J, Yang ZX, Liu Y, Zhang Y et al. Association between the XRCC3 polymorphisms and breast cancer risk: meta-analysis based on case-control studies. Mol Biol Rep 2012; 39: 5125–5134.
Shamseldin HE, Elfaki M, Alkuraya FS . Exome sequencing reveals a novel Fanconi group defined by XRCC2 mutation. J Med Genet 2012; 49: 184–186.
Ludwig T, Chapman DL, Papaioannou VE, Efstratiadis A . Targeted mutations of breast cancer susceptibility gene homologs in mice: lethal phenotypes of Brca1, Brca2, Brca1/Brca2, Brca1/p53, and Brca2/p53 nullizygous embryos. Genes Dev 1997; 11: 1226–1241.
Bahassi EM, Ovesen JL, Riesenberg AL, Bernstein WZ, Hasty PE, Stambrook PJ . The checkpoint kinases Chk1 and Chk2 regulate the functional associations between hBRCA2 and Rad51 in response to DNA damage. Oncogene 2008; 27: 3977–3985.
Sy SM, Huen MS, Chen J . MRG15 is a novel PALB2-interacting factor involved in homologous recombination. J Biol Chem 2009; 284: 21127–21131.
Hayakawa T, Zhang F, Hayakawa N, Ohtani Y, Shinmyozu K, Nakayama J et al. MRG15 binds directly to PALB2 and stimulates homology-directed repair of chromosomal breaks. J Cell Sci 2010; 123: 1124–11230.
Schlacher K, Christ N, Siaud N, Egashira A, Wu H, Jasin M . Double-strand break repair-independent role for BRCA2 in blocking stalled replication fork degradation by MRE11. Cell 2011; 145: 529–542.
Villani G, Hubscher U, Gironis N, Parkkinen S, Pospiech H, Shevelev I et al. In vitro gap-directed translesion DNA synthesis of an abasic site involving human DNA polymerases epsilon, lambda, and beta. J Biol Chem 2011; 286: 32094–32104.
Göhler T, Munoz IM, Rouse J, Blow JJ . PTIP/swift is required for efficient PCNA ubiquitination in response to DNA damage. DNA Repair 2008; 7: 775–787.
Shevelev IV, Ramadan K, Hübscher U . The TREX2 3′->5′ exonuclease physically interacts with DNA polymerase delta and increases its accuracy. ScientificWorld Journal 2002; 2: 275–281.
Maloisel L, Fabre F, Gangloff S . DNA polymerase delta is preferentially recruited during homologous recombination to promote heteroduplex DNA extension. Mol Cell Biol 2008; 28: 1373–1382.
Varadi V, Bevier M, Grzybowska E, Johansson R, Enquist K, Henriksson R et al. Genetic variation in genes encoding for polymerase ζ subunits associates with breast cancer risk, tumour characteristics and survival. Breast Cancer Res Treat 2011; 129: 235–245.
Huertas P . DNA resection in eukaryotes: deciding how to fix the break. Nat Struct Mol Biol 2010; 17: 11–16.
Francis R, Richardson C . Multipotent hematopoietic cells susceptible to alternative double-strand break repair pathways that promote genome rearrangements. Genes Dev 2007; 21: 1064–1074.
Keimling M, Volcic M, Csernok A, Wieland B, Dörk T, Wiesmüller L . Functional characterization connects individual patientś mutations in Ataxia Telangiectasia Mutated (ATM) with dysfunction of specific DNA double-strand break repair signalling pathways. FASEB J 2011; 25: 3849–3860.
Cordelli E, Cinelli S, Lascialfari A, Ranaldi R, Pacchierotti F . Melphalan-induced DNA damage in p53(+/−) and wild type mice analysed by the comet assay. Mutat Res 2004; 550: 133–143.
Gutekunst M, Oren M, Weilbacher A, Dengler MA, Markwardt C, Thomale J et al. p53 hypersensitivity is the predominant mechanism of the unique responsiveness of testicular germ cell tumor (TGCT) cells to cisplatin. PLoS One 2011; 6: e19198.
Tchatchou S, Riedel A, Lyer S, Schmutzhard J, Strobel-Freidekind O, Gronert-Sum S et al. Identification of a DMBT1 polymorphism associated with increased breast cancer risk and decreased promoter activity. Hum Mutat 2010; 31: 60–66.
Ralhan R, Kaur J, Kreienberg R, Wiesmüller L . Links between DNA double strand break repair and breast cancer: accumulating evidence from both familial and nonfamilial cases. Cancer Lett 2007; 248: 1–17.
Toyota M, Suzuki H . Epigenetic drivers of genetic alterations. Adv Genet 2010; 70: 309–323.
Jerry DJ, Kuperwasser C, Downing SR, Pinkas J, He C, Dickinson E et al. Delayed involution of the mammary epithelium in BALB/c-p53null mice. Oncogene 1998; 17: 2305–2312.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003; 13: 2498–2504.
Acknowledgements
We are indebted to Bing Xia, The Cancer Institute of New Jersey UMDNJ Robert Wood Johnson Medical School, New Brunswick, NJ, USA, for the generous gift of EUFA1341 cells and expression plasmid for human PALB2, POZ-PALB2. Jeffrey Kane, Amherst, provided assistance with the cloning of mouse Palb2. We cordially thank Marlen Keimling, Ulm, for experimental help during the initial phase of the project and Richard Friedl, Würzburg, for expert flow cytometry. The drugs KU-55933 and NU7441 were kindly provided by KuDOS Pharmaceuticals Ltd. This work was supported by grants to LW by the Foundation of the State Baden-Württemberg Germany (10-1907-Wi 2), the Ministry for Education and Research, Germany (BMBF, 012P0505), and the German Research Foundation (DFG, WI 3099/7-1 and -2), as well as to DJJ by the National Cancer Institute (R01CA105452).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Böhringer, M., Obermeier, K., Griner, N. et al. siRNA screening identifies differences in the Fanconi anemia pathway in BALB/c-Trp53+/− with susceptibility versus C57BL/6-Trp53+/− mice with resistance to mammary tumors. Oncogene 32, 5458–5470 (2013). https://doi.org/10.1038/onc.2013.38
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.38
Keywords
This article is cited by
-
Genetic modifiers regulating DNA replication and double-strand break repair are associated with differences in mammary tumors in mouse models of Li-Fraumeni syndrome
Oncogene (2021)
-
Heterozygous PALB2 c.1592delT mutation channels DNA double-strand break repair into error-prone pathways in breast cancer patients
Oncogene (2016)